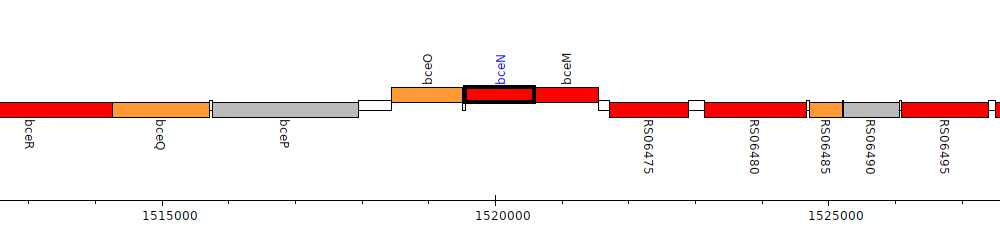
Burkholderia dolosa AU0158, AK34_RS06465 (bceN)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0045226 | extracellular polysaccharide biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: BGD:Bcep1808_4472
|
ECO:0000250 sequence similarity evidence used in manual assertion |
19948863 | Reviewed by curator |
| Molecular Function | GO:0008446 | GDP-mannose 4,6-dehydratase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01472
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019673 | GDP-mannose metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01472
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00520 | Amino sugar and nucleotide sugar metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | GDP-L-fucose biosynthesis I (from GDP-D-mannose) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | GDP-mycosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | GDP-6-deoxy-D-talose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | GDP-L-colitose biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | GDP-D-perosamine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00051 | Fructose and mannose metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd05260 | GDP_MD_SDR_e | 7 | 339 | 1.14257E-172 | ||
| Pfam | PF16363 | GDP-mannose 4,6 dehydratase | IPR016040 | NAD(P)-binding domain | 9 | 334 | 5.3E-137 |
| Gene3D | G3DSA:3.40.50.720 | 8 | 325 | 1.1E-147 | |||
| Gene3D | G3DSA:3.90.25.10 | 191 | 339 | 1.1E-147 | |||
| Hamap | MF_00955 | GDP-mannose 4,6-dehydratase [gmd]. | IPR006368 | GDP-mannose 4,6-dehydratase | 6 | 344 | 52.746 |
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 6 | 338 | 2.22E-74 | |
| TIGRFAM | TIGR01472 | gmd: GDP-mannose 4,6-dehydratase | IPR006368 | GDP-mannose 4,6-dehydratase | 6 | 340 | 4.4E-144 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.