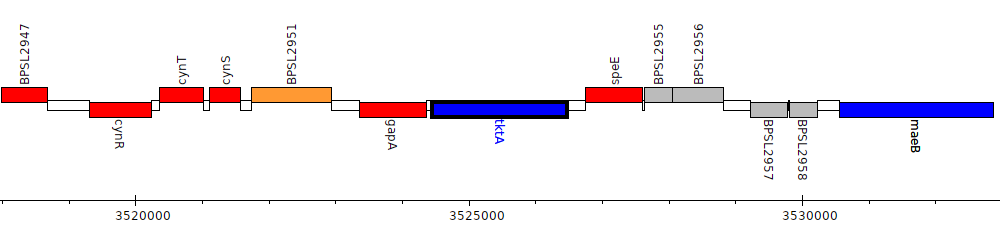
Burkholderia pseudomallei K96243, BPSL2953 (tktA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.50.920
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004802 | transketolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00232
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | thiazole biosynthesis II (aerobic bacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00710 | Carbon fixation in photosynthetic organisms | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | formaldehyde assimilation II (assimilatory RuMP Cycle) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | thiazole biosynthesis I (facultative anaerobic bacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00900 | Terpenoid backbone biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | superpathway of glucose and xylose degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00030 | Pentose phosphate pathway | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | methylerythritol phosphate pathway II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00030 | Pentose phosphate pathway | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00730 | Thiamine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | Rubisco shunt | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bps00710 | Carbon fixation in photosynthetic organisms | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bps01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.920 | IPR009014 | Transketolase C-terminal/Pyruvate-ferredoxin oxidoreductase domain II | 548 | 675 | 6.4E-43 | |
| Gene3D | G3DSA:3.40.50.970 | 331 | 546 | 2.4E-79 | |||
| CDD | cd07033 | TPP_PYR_DXS_TK_like | 363 | 529 | 5.03888E-57 | ||
| SMART | SM00861 | IPR005475 | Transketolase-like, pyrimidine-binding domain | 359 | 534 | 1.7E-63 | |
| TIGRFAM | TIGR00232 | tktlase_bact: transketolase | IPR005478 | Transketolase, bacterial-like | 12 | 674 | 2.9E-292 |
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 9 | 324 | 5.12E-120 | |
| Pfam | PF00456 | Transketolase, thiamine diphosphate binding domain | IPR005474 | Transketolase, N-terminal | 10 | 338 | 1.4E-160 |
| SUPERFAMILY | SSF52922 | IPR009014 | Transketolase C-terminal/Pyruvate-ferredoxin oxidoreductase domain II | 542 | 675 | 3.27E-44 | |
| Gene3D | G3DSA:3.40.50.970 | 6 | 330 | 1.7E-132 | |||
| Pfam | PF02779 | Transketolase, pyrimidine binding domain | IPR005475 | Transketolase-like, pyrimidine-binding domain | 358 | 532 | 1.3E-43 |
| ProSitePatterns | PS00801 | Transketolase signature 1. | IPR005474 | Transketolase, N-terminal | 17 | 37 | - |
| CDD | cd02012 | TPP_TK | IPR005474 | Transketolase, N-terminal | 14 | 278 | 8.79004E-145 |
| Pfam | PF02780 | Transketolase, C-terminal domain | IPR033248 | Transketolase, C-terminal domain | 562 | 666 | 1.7E-17 |
| SUPERFAMILY | SSF52518 | IPR029061 | Thiamin diphosphate-binding fold | 354 | 533 | 3.88E-62 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.