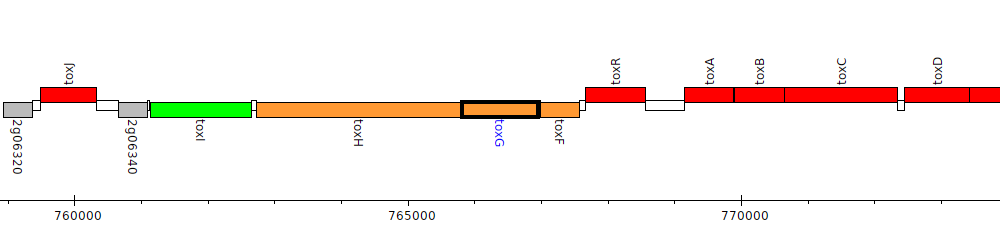
Burkholderia glumae BGR1, bglu_2g06370 (toxG)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01730
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.40.420.20 | 279 | 352 | 2.4E-70 | |||
| TIGRFAM | TIGR01730 | RND_mfp: efflux transporter, RND family, MFP subunit | IPR006143 | RND efflux pump, membrane fusion protein | 43 | 354 | 1.7E-76 |
| Gene3D | G3DSA:2.40.50.100 | 67 | 195 | 2.4E-70 | |||
| Gene3D | G3DSA:1.10.287.470 | 102 | 161 | 2.4E-70 | |||
| Pfam | PF16576 | Barrel-sandwich domain of CusB or HlyD membrane-fusion | IPR032317 | RND efflux pump, membrane fusion protein, barrel-sandwich domain | 67 | 268 | 5.7E-30 |
| Gene3D | G3DSA:2.40.30.170 | 55 | 278 | 2.4E-70 | |||
| SUPERFAMILY | SSF111369 | 56 | 273 | 1.83E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.