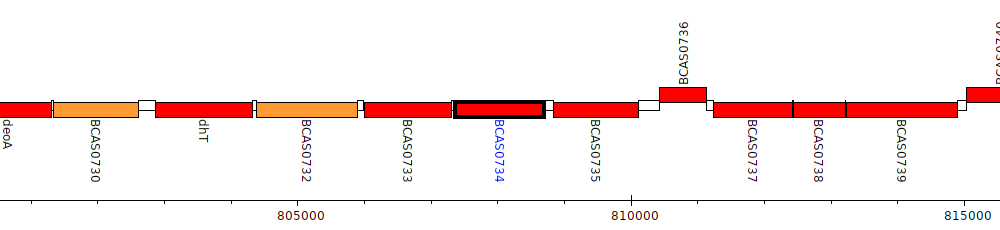
Burkholderia cenocepacia J2315, BCAS0734
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46548
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00910 | Nitrogen metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00250 | Alanine, aspartate and glutamate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF14691 | Dihydroprymidine dehydrogenase domain II, 4Fe-4S cluster | IPR028261 | Dihydroprymidine dehydrogenase domain II | 22 | 130 | 5.1E-36 |
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 143 | 426 | 6.1E-38 |
| PRINTS | PR00419 | Adrenodoxin reductase family signature | 166 | 179 | 7.5E-19 | ||
| PRINTS | PR00419 | Adrenodoxin reductase family signature | 209 | 219 | 7.5E-19 | ||
| PRINTS | PR00419 | Adrenodoxin reductase family signature | 143 | 165 | 7.5E-19 | ||
| SUPERFAMILY | SSF46548 | IPR009051 | Alpha-helical ferredoxin | 11 | 152 | 4.92E-31 | |
| SUPERFAMILY | SSF51971 | 140 | 372 | 5.18E-57 | |||
| Gene3D | G3DSA:1.10.1060.10 | IPR009051 | Alpha-helical ferredoxin | 7 | 143 | 2.1E-40 | |
| Gene3D | G3DSA:3.50.50.60 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 145 | 433 | 1.5E-50 | |
| PRINTS | PR00419 | Adrenodoxin reductase family signature | 278 | 292 | 7.5E-19 | ||
| Gene3D | G3DSA:3.50.50.60 | IPR036188 | FAD/NAD(P)-binding domain superfamily | 241 | 371 | 1.5E-50 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.