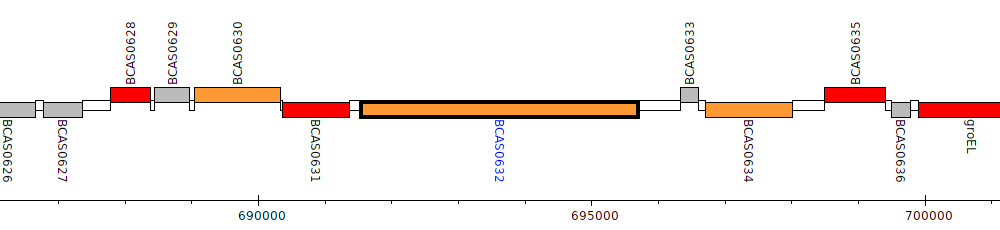
Burkholderia cenocepacia J2315, BCAS0632
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:PF00512
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50123
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:PS50122
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:PS50110
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00512
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000156 | phosphorelay response regulator activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50122
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008984 | protein-glutamate methylesterase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS50122
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:PS50122
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj02030 | Bacterial chemotaxis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj02020 | Two-component system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00388 | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 1019 | 1087 | 7.0E-13 | |
| PRINTS | PR00996 | Glutamate methyltransferase family signature | IPR000780 | MCP methyltransferase, CheR-type | 257 | 270 | 1.0E-27 |
| Gene3D | G3DSA:3.40.50.12740 | 1262 | 1379 | 1.1E-32 | |||
| CDD | cd00156 | REC | IPR001789 | Signal transduction response regulator, receiver domain | 1266 | 1376 | 1.97069E-27 |
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 1265 | 1376 | 2.8E-21 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 889 | 989 | 3.11328E-12 |
| Coils | Coil | 675 | 755 | - | |||
| SUPERFAMILY | SSF55785 | IPR035965 | PAS domain superfamily | 770 | 872 | 6.46E-7 | |
| ProSiteProfiles | PS50110 | Response regulatory domain profile. | IPR001789 | Signal transduction response regulator, receiver domain | 1264 | 1380 | 35.361 |
| SUPERFAMILY | SSF47757 | 233 | 303 | 2.62E-11 | |||
| SUPERFAMILY | SSF52172 | IPR011006 | CheY-like superfamily | 1262 | 1377 | 1.22E-31 | |
| PRINTS | PR00996 | Glutamate methyltransferase family signature | IPR000780 | MCP methyltransferase, CheR-type | 334 | 354 | 1.0E-27 |
| SUPERFAMILY | SSF52738 | IPR035909 | Methylesterase CheB, C-terminal | 24 | 208 | 6.93E-49 | |
| Pfam | PF13596 | PAS domain | 759 | 864 | 7.3E-24 | ||
| PRINTS | PR00996 | Glutamate methyltransferase family signature | IPR000780 | MCP methyltransferase, CheR-type | 370 | 386 | 1.0E-27 |
| Pfam | PF01339 | CheB methylesterase | IPR000673 | Signal transduction response regulator, chemotaxis, protein-glutamate methylesterase | 27 | 202 | 6.3E-48 |
| SUPERFAMILY | SSF55874 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 1076 | 1244 | 5.11E-42 | |
| CDD | cd02440 | AdoMet_MTases | 334 | 472 | 9.68724E-5 | ||
| SUPERFAMILY | SSF55785 | IPR035965 | PAS domain superfamily | 863 | 990 | 1.06E-24 | |
| Gene3D | G3DSA:3.30.450.20 | 742 | 877 | 3.7E-13 | |||
| PRINTS | PR00996 | Glutamate methyltransferase family signature | IPR000780 | MCP methyltransferase, CheR-type | 455 | 478 | 1.0E-27 |
| ProSiteProfiles | PS50109 | Histidine kinase domain profile. | IPR005467 | Histidine kinase domain | 1026 | 1244 | 46.014 |
| Gene3D | G3DSA:3.30.565.10 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 1089 | 1248 | 3.0E-38 | |
| SUPERFAMILY | SSF55785 | IPR035965 | PAS domain superfamily | 546 | 653 | 2.42E-5 | |
| SMART | SM00448 | IPR001789 | Signal transduction response regulator, receiver domain | 1263 | 1376 | 9.2E-31 | |
| ProSiteProfiles | PS50122 | CheB-type methylesterase domain profile. | IPR000673 | Signal transduction response regulator, chemotaxis, protein-glutamate methylesterase | 26 | 204 | 22.08 |
| Gene3D | G3DSA:3.40.50.150 | 300 | 498 | 3.2E-62 | |||
| Pfam | PF13426 | PAS domain | IPR000014 | PAS domain | 890 | 990 | 8.7E-12 |
| Pfam | PF03705 | CheR methyltransferase, all-alpha domain | IPR022641 | Chemotaxis receptor methyltransferase CheR, N-terminal | 238 | 286 | 3.5E-11 |
| SMART | SM00091 | IPR000014 | PAS domain | 757 | 822 | 52.0 | |
| ProSiteProfiles | PS50123 | CheR-type methyltransferase domain profile. | IPR000780 | MCP methyltransferase, CheR-type | 245 | 477 | 55.66 |
| CDD | cd16434 | CheB-CheR_fusion | 26 | 205 | 1.72299E-79 | ||
| SUPERFAMILY | SSF53335 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase | 301 | 495 | 9.46E-49 | |
| Gene3D | G3DSA:3.30.450.20 | 889 | 1000 | 1.4E-22 | |||
| SMART | SM00091 | IPR000014 | PAS domain | 878 | 945 | 2.0E-9 | |
| ProSiteProfiles | PS50113 | PAC domain profile. | IPR000700 | PAS-associated, C-terminal | 825 | 875 | 8.966 |
| SMART | SM00387 | IPR003594 | Histidine kinase/HSP90-like ATPase | 1133 | 1244 | 5.4E-34 | |
| Gene3D | G3DSA:1.10.287.130 | 1006 | 1088 | 6.2E-17 | |||
| Gene3D | G3DSA:3.40.50.180 | IPR035909 | Methylesterase CheB, C-terminal | 18 | 213 | 1.3E-56 | |
| Pfam | PF01739 | CheR methyltransferase, SAM binding domain | IPR022642 | MCP methyltransferase, CheR-type, SAM-binding domain, C-terminal | 302 | 491 | 1.3E-52 |
| SUPERFAMILY | SSF47384 | IPR036097 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain superfamily | 1008 | 1088 | 2.75E-16 | |
| Pfam | PF00512 | His Kinase A (phospho-acceptor) domain | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 1020 | 1087 | 8.4E-11 |
| CDD | cd00082 | HisKA | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 1017 | 1083 | 1.9958E-11 |
| PRINTS | PR00996 | Glutamate methyltransferase family signature | IPR000780 | MCP methyltransferase, CheR-type | 436 | 450 | 1.0E-27 |
| ProSiteProfiles | PS50112 | PAS repeat profile. | IPR000014 | PAS domain | 876 | 947 | 16.574 |
| TIGRFAM | TIGR00229 | sensory_box: PAS domain S-box protein | IPR000014 | PAS domain | 876 | 995 | 7.7E-23 |
| PRINTS | PR00996 | Glutamate methyltransferase family signature | IPR000780 | MCP methyltransferase, CheR-type | 294 | 306 | 1.0E-27 |
| SMART | SM00138 | IPR000780 | MCP methyltransferase, CheR-type | 235 | 496 | 1.3E-83 | |
| SMART | SM00091 | IPR000014 | PAS domain | 546 | 611 | 0.44 | |
| CDD | cd00075 | HATPase | 1139 | 1240 | 1.51933E-31 | ||
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 1134 | 1242 | 2.2E-24 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.