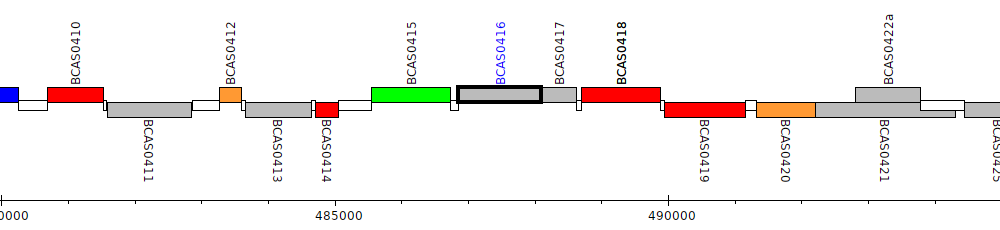
Burkholderia cenocepacia J2315, BCAS0416
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0042128 | nitrate assimilation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00174
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030151 | molybdenum ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF03404
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00407
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00407
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00920 | Sulfur metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00407 | Eukaryotic molybdopterin domain signature | IPR008335 | Eukaryotic molybdopterin oxidoreductase | 96 | 112 | 6.1E-20 |
| PRINTS | PR00407 | Eukaryotic molybdopterin domain signature | IPR008335 | Eukaryotic molybdopterin oxidoreductase | 252 | 265 | 6.1E-20 |
| Pfam | PF03404 | Mo-co oxidoreductase dimerisation domain | IPR005066 | Moybdenum cofactor oxidoreductase, dimerisation | 289 | 402 | 2.1E-16 |
| PRINTS | PR00407 | Eukaryotic molybdopterin domain signature | IPR008335 | Eukaryotic molybdopterin oxidoreductase | 148 | 159 | 6.1E-20 |
| Gene3D | G3DSA:2.60.40.650 | 293 | 418 | 4.6E-46 | |||
| SUPERFAMILY | SSF81296 | IPR014756 | Immunoglobulin E-set | 293 | 409 | 1.82E-29 | |
| PRINTS | PR00407 | Eukaryotic molybdopterin domain signature | IPR008335 | Eukaryotic molybdopterin oxidoreductase | 173 | 188 | 6.1E-20 |
| SUPERFAMILY | SSF56524 | IPR036374 | Oxidoreductase, molybdopterin-binding domain superfamily | 59 | 298 | 1.03E-70 | |
| TIGRFAM | TIGR04555 | sulfite_DH_soxC: sulfite dehydrogenase | IPR030835 | Sulfite dehydrogenase SoxC | 15 | 418 | 4.2E-182 |
| PRINTS | PR00407 | Eukaryotic molybdopterin domain signature | IPR008335 | Eukaryotic molybdopterin oxidoreductase | 236 | 249 | 6.1E-20 |
| PRINTS | PR00407 | Eukaryotic molybdopterin domain signature | IPR008335 | Eukaryotic molybdopterin oxidoreductase | 374 | 386 | 6.1E-20 |
| Pfam | PF00174 | Oxidoreductase molybdopterin binding domain | IPR000572 | Oxidoreductase, molybdopterin-binding domain | 108 | 268 | 8.8E-48 |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 38 | 8.041 |
| Gene3D | G3DSA:3.90.420.10 | IPR036374 | Oxidoreductase, molybdopterin-binding domain superfamily | 38 | 292 | 2.3E-81 | |
| PRINTS | PR00407 | Eukaryotic molybdopterin domain signature | IPR008335 | Eukaryotic molybdopterin oxidoreductase | 317 | 331 | 6.1E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.