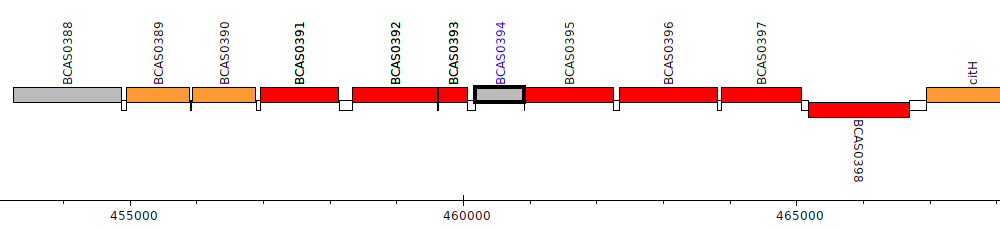
Burkholderia cenocepacia J2315, BCAS0394
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | allantoin degradation to glyoxylate II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | allantoin degradation to ureidoglycolate II (ammonia producing) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF05899 | Protein of unknown function (DUF861) | IPR008579 | Domain of unknown function DUF861, cupin-3 | 166 | 232 | 5.9E-9 |
| SUPERFAMILY | SSF51182 | IPR011051 | RmlC-like cupin domain superfamily | 137 | 234 | 2.22E-14 | |
| Gene3D | G3DSA:2.60.120.10 | IPR014710 | RmlC-like jelly roll fold | 26 | 130 | 3.6E-70 | |
| Gene3D | G3DSA:2.60.120.10 | IPR014710 | RmlC-like jelly roll fold | 131 | 233 | 3.6E-70 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.