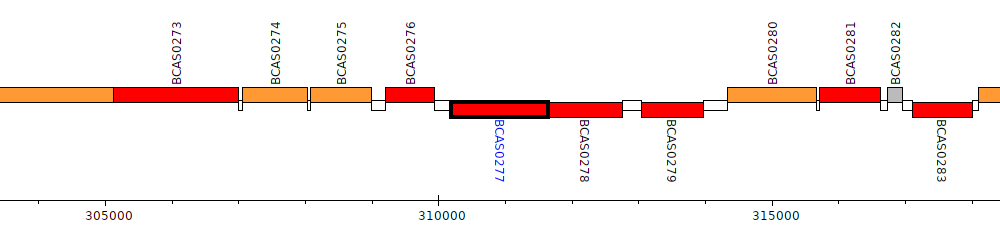
Burkholderia cenocepacia J2315, BCAS0277
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.309.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.309.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.309.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00340 | Histidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00620 | Pyruvate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00040 | Pentose and glucuronate interconversions | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00280 | Valine, leucine and isoleucine degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00410 | beta-Alanine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00010 | Glycolysis / Gluconeogenesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00903 | Limonene and pinene degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00310 | Lysine degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00625 | Chloroalkane and chloroalkene degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00071 | Fatty acid degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00561 | Glycerolipid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00330 | Arginine and proline metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00380 | Tryptophan metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00053 | Ascorbate and aldarate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.605.10 | IPR016162 | Aldehyde dehydrogenase, N-terminal | 21 | 472 | 2.8E-167 | |
| CDD | cd07109 | ALDH_AAS00426 | 22 | 479 | 0.0 | ||
| ProSitePatterns | PS00070 | Aldehyde dehydrogenases cysteine active site. | IPR016160 | Aldehyde dehydrogenase, cysteine active site | 278 | 289 | - |
| Gene3D | G3DSA:3.40.309.10 | IPR016163 | Aldehyde dehydrogenase, C-terminal | 254 | 446 | 2.8E-167 | |
| ProSitePatterns | PS00687 | Aldehyde dehydrogenases glutamic acid active site. | IPR029510 | Aldehyde dehydrogenase, glutamic acid active site | 250 | 257 | - |
| Pfam | PF00171 | Aldehyde dehydrogenase family | IPR015590 | Aldehyde dehydrogenase domain | 16 | 477 | 5.2E-155 |
| SUPERFAMILY | SSF53720 | IPR016161 | Aldehyde/histidinol dehydrogenase | 2 | 480 | 1.96E-152 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.