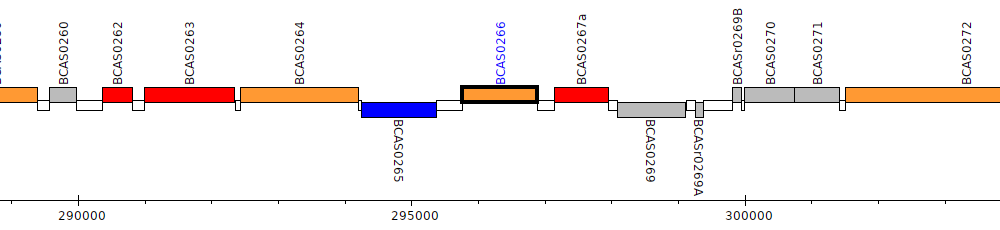
Burkholderia cenocepacia J2315, BCAS0266
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47384
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47384
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| ProSiteProfiles | PS50109 | Histidine kinase domain profile. | IPR005467 | Histidine kinase domain | 158 | 373 | 37.659 |
| Gene3D | G3DSA:3.30.565.10 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 222 | 371 | 1.6E-31 | |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 315 | 325 | 9.3E-8 |
| CDD | cd00075 | HATPase | 265 | 369 | 2.43079E-26 | ||
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 297 | 311 | 9.3E-8 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 334 | 352 | 9.3E-8 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 357 | 370 | 9.3E-8 |
| SUPERFAMILY | SSF47384 | IPR036097 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain superfamily | 147 | 214 | 1.45E-11 | |
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 261 | 371 | 3.7E-17 |
| CDD | cd00082 | HisKA | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 149 | 212 | 9.72042E-9 |
| SMART | SM00387 | IPR003594 | Histidine kinase/HSP90-like ATPase | 260 | 373 | 2.1E-25 | |
| SMART | SM00388 | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 151 | 216 | 8.8E-11 | |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 52 | 80 | - | ||
| Pfam | PF14361 | RsbT co-antagonist protein rsbRD N-terminal domain | IPR025751 | RsbT co-antagonist protein RsbRD, N-terminal domain | 3 | 110 | 5.0E-7 |
| Pfam | PF00512 | His Kinase A (phospho-acceptor) domain | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 152 | 213 | 7.3E-9 |
| Gene3D | G3DSA:1.10.287.130 | 145 | 217 | 8.9E-11 | |||
| SUPERFAMILY | SSF55874 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 205 | 371 | 2.38E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.