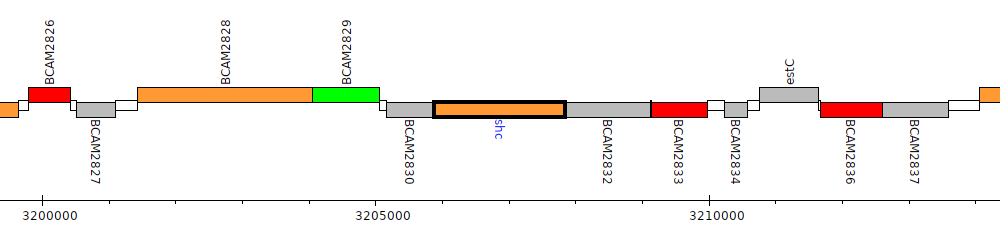
Burkholderia cenocepacia J2315, BCAM2831 (shc)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0005811 | lipid droplet |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01787
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016866 | intramolecular transferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS01074
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0042300 | beta-amyrin synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01787
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000250 | lanosterol synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01787
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019746 | hopanoid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01507
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016104 | triterpenoid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01787
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | diploterol biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00909 | Sesquiterpenoid and triterpenoid biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00909 | Sesquiterpenoid and triterpenoid biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | hopanoid biosynthesis (bacteria) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF48239 | IPR008930 | Terpenoid cyclases/protein prenyltransferase alpha-alpha toroid | 318 | 644 | 6.75E-109 | |
| Pfam | PF13249 | Squalene-hopene cyclase N-terminal domain | IPR032697 | Squalene cyclase, N-terminal | 23 | 313 | 2.2E-115 |
| CDD | cd02892 | SQCY_1 | IPR018333 | Squalene cyclase | 23 | 642 | 0.0 |
| TIGRFAM | TIGR01787 | squalene_cyclas: squalene/oxidosqualene cyclases | IPR018333 | Squalene cyclase | 22 | 644 | 7.7E-188 |
| Gene3D | G3DSA:1.50.10.20 | 44 | 321 | 6.6E-230 | |||
| Pfam | PF13243 | Squalene-hopene cyclase C-terminal domain | IPR032696 | Squalene cyclase, C-terminal | 322 | 644 | 9.2E-149 |
| SUPERFAMILY | SSF48239 | IPR008930 | Terpenoid cyclases/protein prenyltransferase alpha-alpha toroid | 48 | 318 | 2.2E-76 | |
| Gene3D | G3DSA:1.50.10.20 | 25 | 642 | 6.6E-230 | |||
| SFLD | SFLDG01016 | Prenyltransferase Like 2 | 17 | 647 | 0.0 | ||
| TIGRFAM | TIGR01507 | hopene_cyclase: squalene-hopene cyclase | IPR006400 | Squalene hopene cyclase | 19 | 647 | 4.0E-231 |
| ProSitePatterns | PS01074 | Terpene synthases signature. | IPR002365 | Terpene synthase, conserved site | 499 | 513 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.