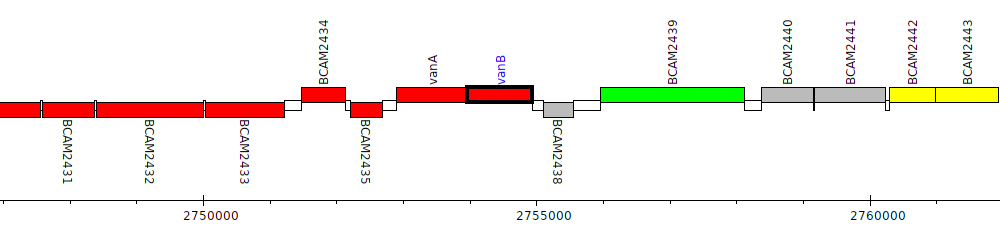
Burkholderia cenocepacia J2315, BCAM2437 (vanB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS51085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PS51384
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051537 | 2 iron, 2 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00197
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS51085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PS51384
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51384
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051537 | 2 iron, 2 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00197
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51384
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00627 | Aminobenzoate degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.10.20.30 | IPR012675 | Beta-grasp domain superfamily | 227 | 321 | 3.4E-24 | |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 52 | 66 | 1.4E-53 |
| Pfam | PF00111 | 2Fe-2S iron-sulfur cluster binding domain | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 242 | 311 | 8.5E-11 |
| CDD | cd06185 | PDR_like | 11 | 224 | 4.19081E-119 | ||
| Pfam | PF00970 | Oxidoreductase FAD-binding domain | IPR008333 | Flavoprotein pyridine nucleotide cytochrome reductase-like, FAD-binding domain | 10 | 102 | 1.6E-4 |
| Gene3D | G3DSA:2.40.30.10 | 1 | 101 | 5.0E-33 | |||
| Gene3D | G3DSA:3.40.50.80 | IPR039261 | Ferredoxin-NADP reductase (FNR), nucleotide-binding domain | 102 | 223 | 9.8E-30 | |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 191 | 199 | 1.4E-53 |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 30 | 40 | 1.4E-53 |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 137 | 146 | 1.4E-53 |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 76 | 90 | 1.4E-53 |
| SUPERFAMILY | SSF63380 | IPR017938 | Riboflavin synthase-like beta-barrel | 5 | 100 | 1.97E-30 | |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 90 | 104 | 1.4E-53 |
| ProSitePatterns | PS00197 | 2Fe-2S ferredoxin-type iron-sulfur binding region signature. | IPR006058 | 2Fe-2S ferredoxin, iron-sulphur binding site | 270 | 278 | - |
| SUPERFAMILY | SSF52343 | IPR039261 | Ferredoxin-NADP reductase (FNR), nucleotide-binding domain | 98 | 228 | 1.44E-32 | |
| ProSiteProfiles | PS51384 | Ferredoxin reductase-type FAD binding domain profile. | IPR017927 | FAD-binding domain, ferredoxin reductase-type | 3 | 105 | 13.764 |
| SUPERFAMILY | SSF54292 | IPR036010 | 2Fe-2S ferredoxin-like superfamily | 213 | 320 | 1.44E-28 | |
| ProSiteProfiles | PS51085 | 2Fe-2S ferredoxin-type iron-sulfur binding domain profile. | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 236 | 321 | 9.972 |
| CDD | cd00207 | fer2 | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 237 | 318 | 3.53154E-18 |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 112 | 131 | 1.4E-53 |
| Pfam | PF00175 | Oxidoreductase NAD-binding domain | IPR001433 | Oxidoreductase FAD/NAD(P)-binding | 113 | 204 | 1.6E-6 |
| PRINTS | PR00409 | Phthalate dioxygenase reductase family signature | IPR000951 | Phthalate dioxygenase reductase | 40 | 52 | 1.4E-53 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.