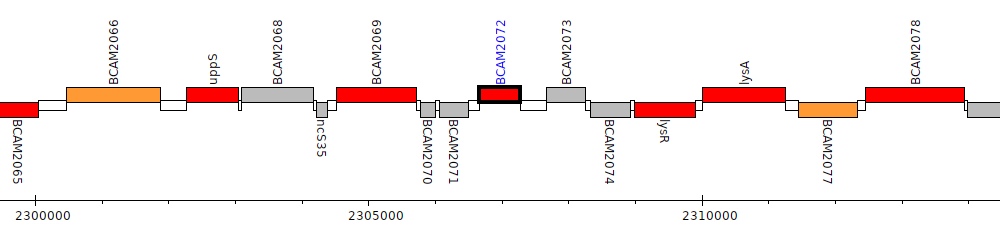
Burkholderia cenocepacia J2315, BCAM2072
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00630 | Glyoxylate and dicarboxylate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.150.240 | IPR023198 | Phosphoglycolate phosphatase-like, domain 2 | 18 | 80 | 5.3E-30 | |
| SUPERFAMILY | SSF56784 | IPR036412 | HAD-like superfamily | 1 | 205 | 5.12E-39 | |
| Gene3D | G3DSA:3.40.50.1000 | IPR023214 | HAD superfamily | 6 | 193 | 5.3E-30 | |
| Pfam | PF13419 | Haloacid dehalogenase-like hydrolase | IPR041492 | Haloacid dehalogenase-like hydrolase | 6 | 177 | 2.9E-22 |
| SFLD | SFLDS00003 | Haloacid Dehalogenase | 2 | 205 | 1.0E-18 | ||
| SFLD | SFLDG01129 | C1.5: HAD, Beta-PGM, Phosphatase Like | 2 | 205 | 1.0E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.