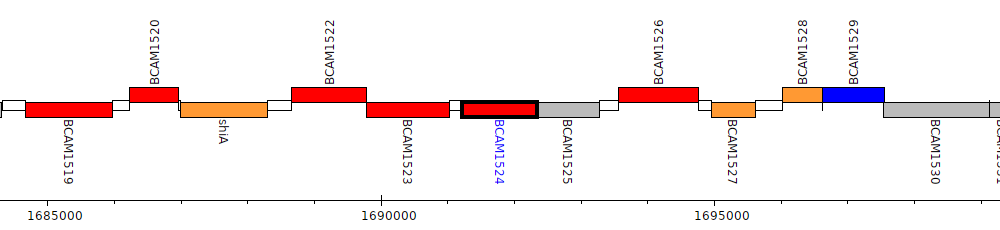
Burkholderia cenocepacia J2315, BCAM1524
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008686 | 3,4-dihydroxy-2-butanone-4-phosphate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00506
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009231 | riboflavin biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00506
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | 6-hydroxymethyl-dihydropterin diphosphate biosynthesis III (Chlamydia) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | flavin biosynthesis II (archaea) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | flavin biosynthesis III (fungi) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | toxoflavin biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00740 | Riboflavin metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00740 | Riboflavin metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00790 | Folate biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF001259 | 305 | 363 | 8.3E-6 | |||
| Gene3D | G3DSA:3.40.50.10990 | IPR036144 | RibA-like superfamily | 207 | 356 | 6.4E-22 | |
| PIRSF | PIRSF001259 | 3 | 301 | 1.3E-101 | |||
| Gene3D | G3DSA:3.90.870.10 | 3 | 206 | 2.9E-86 | |||
| Pfam | PF00925 | GTP cyclohydrolase II | IPR032677 | GTP cyclohydrolase II | 208 | 298 | 2.4E-16 |
| TIGRFAM | TIGR00506 | ribB: 3,4-dihydroxy-2-butanone-4-phosphate synthase | IPR000422 | 3,4-dihydroxy-2-butanone 4-phosphate synthase, RibB | 9 | 199 | 1.6E-70 |
| Pfam | PF00926 | 3,4-dihydroxy-2-butanone 4-phosphate synthase | IPR000422 | 3,4-dihydroxy-2-butanone 4-phosphate synthase, RibB | 8 | 198 | 1.1E-81 |
| Pfam | PF00925 | GTP cyclohydrolase II | IPR032677 | GTP cyclohydrolase II | 309 | 353 | 3.6E-5 |
| SUPERFAMILY | SSF142695 | IPR036144 | RibA-like superfamily | 204 | 355 | 2.35E-37 | |
| Hamap | MF_00180 | 3,4-dihydroxy-2-butanone 4-phosphate synthase [ribB]. | IPR000422 | 3,4-dihydroxy-2-butanone 4-phosphate synthase, RibB | 1 | 203 | 38.92 |
| SUPERFAMILY | SSF55821 | IPR017945 | DHBP synthase RibB-like alpha/beta domain superfamily | 5 | 203 | 1.81E-84 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.