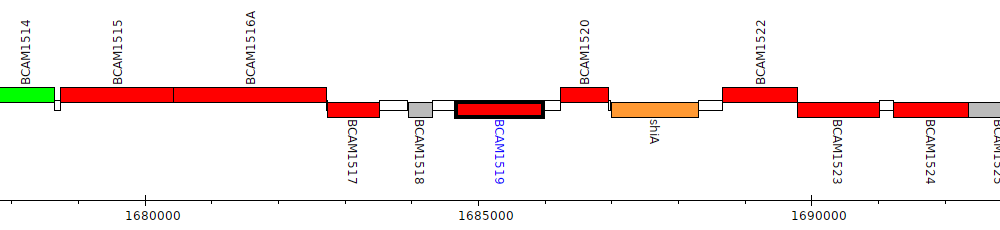
Burkholderia cenocepacia J2315, BCAM1519
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55205
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF00275
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00520 | Amino sugar and nucleotide sugar metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00550 | Peptidoglycan biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.65.10.10 | IPR036968 | Enolpyruvate transferase domain superfamily | 21 | 232 | 4.0E-108 | |
| Pfam | PF00275 | EPSP synthase (3-phosphoshikimate 1-carboxyvinyltransferase) | IPR001986 | Enolpyruvate transferase domain | 7 | 420 | 1.3E-49 |
| SUPERFAMILY | SSF55205 | IPR013792 | RNA 3'-terminal phosphate cyclase/enolpyruvate transferase, alpha/beta | 1 | 431 | 1.64E-88 | |
| Gene3D | G3DSA:3.65.10.10 | IPR036968 | Enolpyruvate transferase domain superfamily | 6 | 427 | 4.0E-108 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.