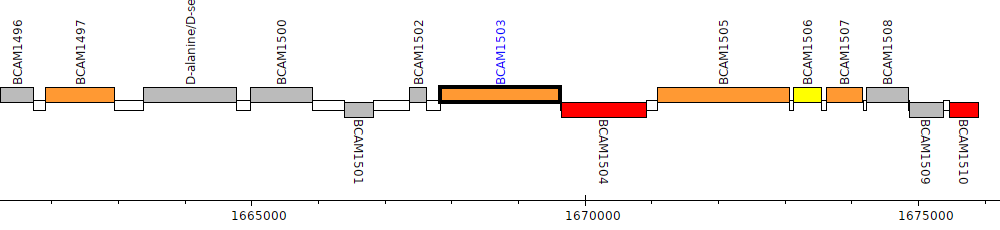
Burkholderia cenocepacia J2315, BCAM1503
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS50885
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:PS50111
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS50111
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj02020 | Two-component system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj02030 | Bacterial chemotaxis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00672 | HAMP domain | IPR003660 | HAMP domain | 297 | 345 | 3.7E-12 |
| Coils | Coil | 554 | 574 | - | |||
| CDD | cd11386 | MCP_signal | 377 | 576 | 1.05802E-54 | ||
| Gene3D | G3DSA:3.30.450.20 | 55 | 267 | 5.1E-52 | |||
| Pfam | PF00015 | Methyl-accepting chemotaxis protein (MCP) signalling domain | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 410 | 564 | 2.4E-51 |
| SMART | SM00304 | IPR003660 | HAMP domain | 295 | 349 | 3.8E-13 | |
| SMART | SM00283 | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 350 | 597 | 2.6E-89 | |
| ProSiteProfiles | PS50111 | Bacterial chemotaxis sensory transducers domain profile. | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 354 | 583 | 50.798 |
| CDD | cd12912 | PDC2_MCP_like | 177 | 263 | 4.74557E-11 | ||
| CDD | cd06225 | HAMP | IPR003660 | HAMP domain | 299 | 346 | 1.72433E-9 |
| ProSiteProfiles | PS50885 | HAMP domain profile. | IPR003660 | HAMP domain | 295 | 349 | 10.849 |
| CDD | cd12913 | PDC1_MCP_like | 95 | 170 | 2.56065E-22 | ||
| Gene3D | G3DSA:3.30.450.20 | 64 | 168 | 5.1E-52 | |||
| SUPERFAMILY | SSF58104 | 317 | 598 | 2.09E-71 | |||
| Pfam | PF02743 | Cache domain | IPR033479 | Double Cache domain 1 | 26 | 257 | 1.1E-30 |
| SUPERFAMILY | SSF103190 | IPR029151 | Periplasmic sensor-like domain superfamily | 107 | 177 | 7.32E-14 | |
| Gene3D | G3DSA:1.10.287.950 | 295 | 599 | 1.7E-84 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.