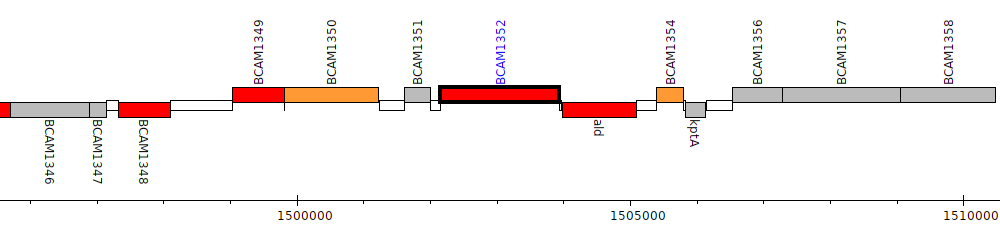
Burkholderia cenocepacia J2315, BCAM1352
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003887 | DNA-directed DNA polymerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00483
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR00870
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF89550
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:PR00870
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.150.20 | 89 | 152 | 5.1E-19 | |||
| Gene3D | G3DSA:1.10.150.110 | IPR027421 | DNA polymerase lambda lyase domain superfamily | 2 | 87 | 5.3E-22 | |
| SUPERFAMILY | SSF47802 | IPR027421 | DNA polymerase lambda lyase domain superfamily | 2 | 81 | 1.31E-18 | |
| SMART | SM00278 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 94 | 113 | 1.8 | |
| SMART | SM00481 | IPR003141 | Polymerase/histidinol phosphatase, N-terminal | 341 | 419 | 6.5E-9 | |
| Pfam | PF14716 | Helix-hairpin-helix domain | IPR010996 | DNA polymerase beta-like, N-terminal domain | 5 | 74 | 4.8E-20 |
| PRINTS | PR00870 | DNA-polymerase family X pol beta-like signature | IPR002008 | DNA polymerase family X, beta-like | 187 | 206 | 4.8E-8 |
| SMART | SM00278 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 129 | 148 | 350.0 | |
| SMART | SM00278 | IPR003583 | Helix-hairpin-helix DNA-binding motif, class 1 | 54 | 73 | 200.0 | |
| CDD | cd00141 | NT_POLXc | IPR002054 | DNA-directed DNA polymerase X | 6 | 316 | 3.43298E-94 |
| SUPERFAMILY | SSF89550 | IPR016195 | Polymerase/histidinol phosphatase-like | 341 | 571 | 2.62E-40 | |
| SMART | SM00483 | IPR002054 | DNA-directed DNA polymerase X | 3 | 317 | 1.3E-25 | |
| PRINTS | PR00870 | DNA-polymerase family X pol beta-like signature | IPR002008 | DNA polymerase family X, beta-like | 61 | 78 | 4.8E-8 |
| Gene3D | G3DSA:3.30.210.10 | IPR037160 | DNA polymerase, thumb domain superfamily | 264 | 328 | 2.9E-23 | |
| Pfam | PF14520 | Helix-hairpin-helix domain | 95 | 150 | 8.4E-8 | ||
| Gene3D | G3DSA:3.30.460.10 | 159 | 263 | 4.1E-25 | |||
| SUPERFAMILY | SSF158702 | 70 | 149 | 7.06E-7 | |||
| Pfam | PF14791 | DNA polymerase beta thumb | IPR029398 | DNA polymerase beta, thumb domain | 257 | 317 | 2.8E-17 |
| PIRSF | PIRSF005047 | IPR022311 | Uncharacterised conserved protein UCP005047, YshC | 3 | 574 | 2.1E-201 | |
| SUPERFAMILY | SSF81301 | 161 | 317 | 6.54E-39 | |||
| CDD | cd07436 | PHP_PolX | 335 | 570 | 2.6359E-91 | ||
| Gene3D | G3DSA:3.20.20.140 | 329 | 575 | 2.3E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.