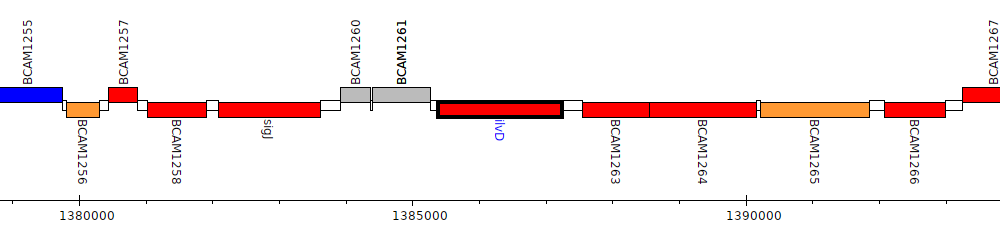
Burkholderia cenocepacia J2315, BCAM1262 (ilvD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00920
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009082 | branched-chain amino acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00012
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004160 | dihydroxy-acid dehydratase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00012
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00290 | Valine, leucine and isoleucine biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00770 | Pantothenate and CoA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-isoleucine biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-isoleucine biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00290 | Valine, leucine and isoleucine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | L-isoleucine biosynthesis IV | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00770 | Pantothenate and CoA biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | pyruvate fermentation to isobutanol (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Coils | Coil | 617 | 619 | - | |||
| ProSitePatterns | PS00886 | Dihydroxy-acid and 6-phosphogluconate dehydratases signature 1. | IPR020558 | Dihydroxy-acid/6-phosphogluconate dehydratase, conserved site | 122 | 132 | - |
| Pfam | PF00920 | Dehydratase family | IPR000581 | Dihydroxy-acid/6-phosphogluconate dehydratase | 34 | 610 | 1.1E-221 |
| ProSitePatterns | PS00887 | Dihydroxy-acid and 6-phosphogluconate dehydratases signature 2. | IPR020558 | Dihydroxy-acid/6-phosphogluconate dehydratase, conserved site | 514 | 525 | - |
| SUPERFAMILY | SSF143975 | IPR037237 | IlvD/EDD, N-terminal domain | 4 | 425 | 1.2E-135 | |
| Gene3D | G3DSA:3.50.30.80 | IPR042096 | Dihydroxy-acid dehydratase, C-terminal | 426 | 578 | 4.0E-55 | |
| TIGRFAM | TIGR00110 | ilvD: dihydroxy-acid dehydratase | IPR004404 | Dihydroxy-acid dehydratase | 18 | 613 | 7.2E-256 |
| SUPERFAMILY | SSF52016 | 426 | 614 | 1.57E-61 | |||
| Hamap | MF_00012 | Dihydroxy-acid dehydratase [ilvD]. | IPR004404 | Dihydroxy-acid dehydratase | 4 | 613 | 44.725 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.