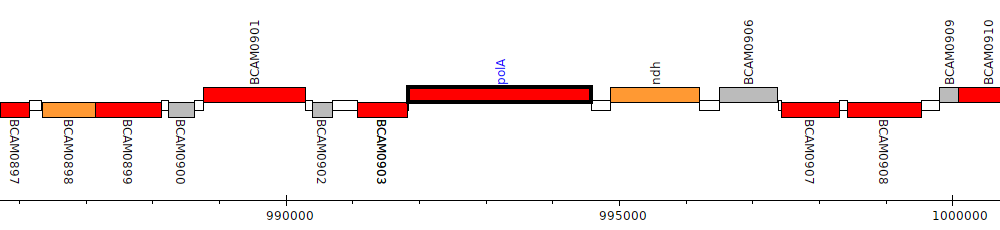
Burkholderia cenocepacia J2315, BCAM0904 (polA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006139 | nucleobase-containing compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01612
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd09898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003887 | DNA-directed DNA polymerase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00482
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006261 | DNA-dependent DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:PR00868
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01612
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008408 | 3'-5' exonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01612
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:SM00482
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd09898
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03440 | Homologous recombination | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03420 | Nucleotide excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03410 | Base excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03030 | DNA replication | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF47807 | IPR036279 | 5'-3' exonuclease, C-terminal domain superfamily | 174 | 292 | 6.73E-28 | |
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 689 | 712 | 9.2E-76 |
| Gene3D | G3DSA:3.40.50.1010 | 12 | 262 | 2.7E-85 | |||
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 780 | 791 | 9.2E-76 |
| Gene3D | G3DSA:1.10.150.20 | 177 | 249 | 2.7E-85 | |||
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 864 | 877 | 9.2E-76 |
| Pfam | PF01367 | 5'-3' exonuclease, C-terminal SAM fold | IPR020045 | DNA polymerase I-like, H3TH domain | 174 | 262 | 4.2E-23 |
| Gene3D | G3DSA:1.10.150.20 | 698 | 841 | 2.2E-108 | |||
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 834 | 850 | 9.2E-76 |
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 644 | 666 | 9.2E-76 |
| Gene3D | G3DSA:3.30.420.10 | IPR036397 | Ribonuclease H superfamily | 317 | 531 | 2.6E-66 | |
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 743 | 768 | 9.2E-76 |
| Pfam | PF02739 | 5'-3' exonuclease, N-terminal resolvase-like domain | IPR020046 | 5'-3' exonuclease, alpha-helical arch, N-terminal | 12 | 173 | 3.0E-50 |
| CDD | cd08637 | DNA_pol_A_pol_I_C | 537 | 913 | 0.0 | ||
| CDD | cd06139 | DNA_polA_I_Ecoli_like_exo | 335 | 526 | 9.22061E-79 | ||
| Gene3D | G3DSA:3.30.70.370 | 653 | 913 | 2.2E-108 | |||
| Pfam | PF01612 | 3'-5' exonuclease | IPR002562 | 3'-5' exonuclease domain | 320 | 505 | 8.4E-33 |
| SMART | SM00475 | IPR002421 | 5'-3' exonuclease, N-terminal | 9 | 264 | 1.9E-109 | |
| SUPERFAMILY | SSF56672 | 511 | 917 | 6.78E-157 | |||
| SUPERFAMILY | SSF53098 | IPR012337 | Ribonuclease H-like superfamily | 323 | 505 | 1.92E-52 | |
| SUPERFAMILY | SSF88723 | IPR029060 | PIN-like domain superfamily | 10 | 173 | 1.46E-53 | |
| Pfam | PF00476 | DNA polymerase family A | IPR001098 | DNA-directed DNA polymerase, family A, palm domain | 540 | 915 | 1.1E-151 |
| CDD | cd09898 | H3TH_53EXO | IPR020045 | DNA polymerase I-like, H3TH domain | 176 | 248 | 1.03053E-27 |
| Gene3D | G3DSA:1.20.1060.10 | 532 | 647 | 2.0E-41 | |||
| TIGRFAM | TIGR00593 | pola: DNA polymerase I | IPR018320 | DNA polymerase 1 | 12 | 917 | 5.0E-272 |
| SMART | SM00482 | IPR001098 | DNA-directed DNA polymerase, family A, palm domain | 675 | 881 | 2.5E-116 | |
| SMART | SM00474 | IPR002562 | 3'-5' exonuclease domain | 320 | 506 | 4.1E-32 | |
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 802 | 813 | 9.2E-76 |
| SMART | SM00279 | IPR008918 | Helix-hairpin-helix motif, class 2 | 176 | 211 | 2.4E-12 | |
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 719 | 732 | 9.2E-76 |
| PRINTS | PR00868 | DNA-polymerase family A (pol I) signature | IPR002298 | DNA polymerase A | 667 | 682 | 9.2E-76 |
| CDD | cd09859 | PIN_53EXO | 14 | 170 | 8.70106E-76 | ||
| ProSitePatterns | PS00447 | DNA polymerase family A signature. | IPR019760 | DNA-directed DNA polymerase, family A, conserved site | 743 | 762 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.