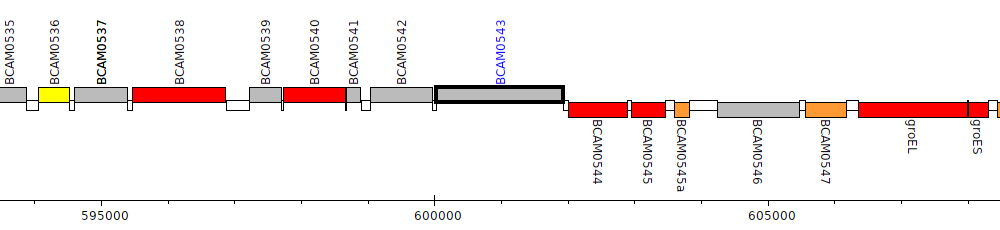
Burkholderia cenocepacia J2315, BCAM0543
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01979
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006534 | cysteine metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01979
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0031071 | cysteine desulfurase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01979
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.640.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00730 | Thiamine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00450 | Selenocompound metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | [2Fe-2S] iron-sulfur cluster biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53383 | IPR015424 | Pyridoxal phosphate-dependent transferase | 236 | 635 | 5.21E-125 | |
| Gene3D | G3DSA:3.90.1150.10 | IPR015422 | Pyridoxal phosphate-dependent transferase domain 1 | 241 | 629 | 1.8E-168 | |
| Gene3D | G3DSA:3.40.640.10 | IPR015421 | Pyridoxal phosphate-dependent transferase, major domain | 269 | 529 | 1.8E-168 | |
| Pfam | PF00266 | Aminotransferase class-V | IPR000192 | Aminotransferase class V domain | 258 | 627 | 9.5E-146 |
| TIGRFAM | TIGR01979 | sufS: cysteine desulfurase, SufS family | IPR010970 | Cysteine desulfurase, SufS | 240 | 633 | 1.5E-170 |
| CDD | cd06453 | SufS_like | 258 | 631 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.