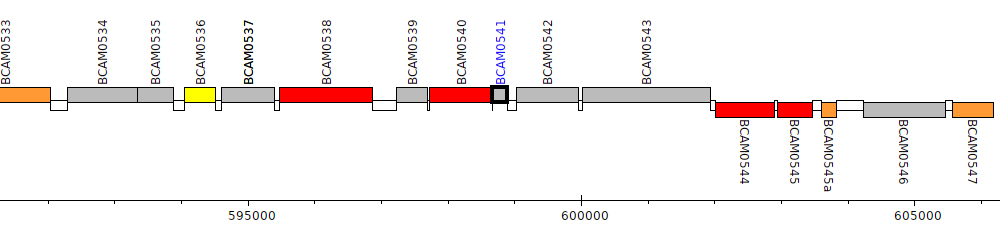
Burkholderia cenocepacia J2315, BCAM0541
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47413
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47413
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00530 | IPR001387 | Cro/C1-type helix-turn-helix domain | 11 | 66 | 7.2E-16 | |
| CDD | cd00093 | HTH_XRE | IPR001387 | Cro/C1-type helix-turn-helix domain | 9 | 66 | 4.54567E-16 |
| ProSiteProfiles | PS50943 | Cro/C1-type HTH domain profile. | IPR001387 | Cro/C1-type helix-turn-helix domain | 12 | 66 | 15.978 |
| SUPERFAMILY | SSF47413 | IPR010982 | Lambda repressor-like, DNA-binding domain superfamily | 2 | 68 | 6.17E-19 | |
| Pfam | PF01381 | Helix-turn-helix | IPR001387 | Cro/C1-type helix-turn-helix domain | 12 | 65 | 2.3E-16 |
| Gene3D | G3DSA:1.10.260.40 | 2 | 71 | 1.9E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.