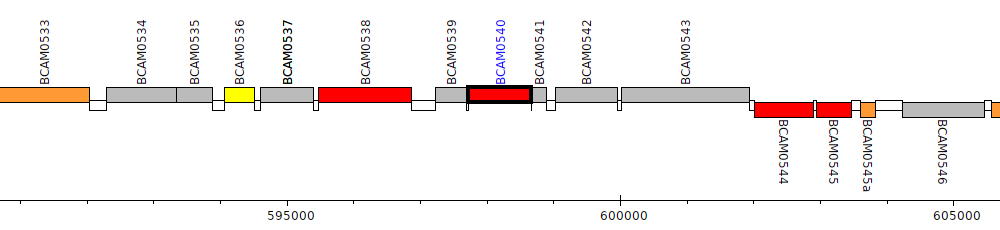
Burkholderia cenocepacia J2315, BCAM0540
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0009001 | serine O-acetyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000441
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006535 | cysteine biosynthetic process from serine |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000441
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000441
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00270 | Cysteine and methionine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00920 | Sulfur metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00920 | Sulfur metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00270 | Cysteine and methionine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | D-cycloserine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | seleno-amino acid biosynthesis (plants) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-cysteine biosynthesis VII (from <i>S</i>-sulfo-L-cysteine) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd03354 | LbH_SAT | 187 | 289 | 8.04273E-46 | ||
| SUPERFAMILY | SSF51161 | IPR011004 | Trimeric LpxA-like superfamily | 110 | 290 | 1.18E-46 | |
| Gene3D | G3DSA:1.10.3130.10 | IPR042122 | Serine acetyltransferase, N-terminal domain superfamily | 33 | 189 | 3.4E-48 | |
| Pfam | PF00132 | Bacterial transferase hexapeptide (six repeats) | IPR001451 | Hexapeptide repeat | 249 | 283 | 2.3E-7 |
| PIRSF | PIRSF000441 | IPR005881 | Serine O-acetyltransferase | 122 | 301 | 3.5E-53 | |
| Gene3D | G3DSA:2.160.10.10 | 190 | 302 | 1.1E-30 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.