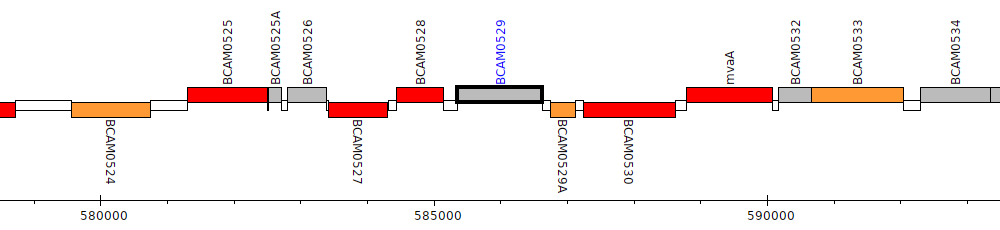
Burkholderia cenocepacia J2315, BCAM0529
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0033961 | cis-stilbene-oxide hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF001112
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00412
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.1820 | IPR029058 | Alpha/Beta hydrolase fold | 37 | 425 | 4.4E-134 | |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 396 | 418 | 2.7E-10 |
| SUPERFAMILY | SSF53474 | IPR029058 | Alpha/Beta hydrolase fold | 36 | 420 | 6.98E-97 | |
| ProSiteProfiles | PS51318 | Twin arginine translocation (Tat) signal profile. | IPR006311 | Twin-arginine translocation pathway, signal sequence | 1 | 38 | 8.107 |
| Pfam | PF06441 | Epoxide hydrolase N terminus | IPR010497 | Epoxide hydrolase, N-terminal | 39 | 143 | 6.2E-35 |
| PIRSF | PIRSF001112 | IPR016292 | Epoxide hydrolase | 7 | 425 | 4.2E-148 | |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 135 | 153 | 2.7E-10 |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 209 | 222 | 2.7E-10 |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 164 | 179 | 2.7E-10 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.