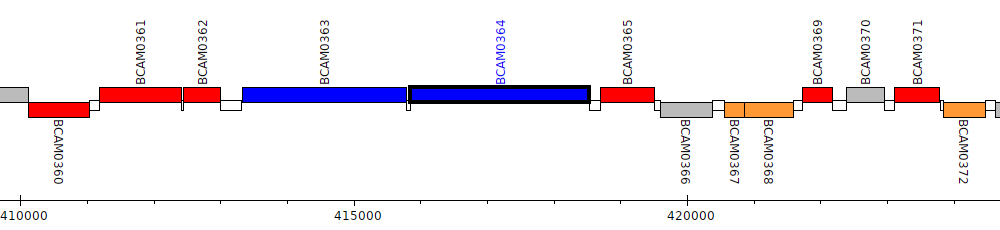
Burkholderia cenocepacia J2315, BCAM0364
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48208
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030246 | carbohydrate binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.70.98.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF07971 | Glycosyl hydrolase family 92 | IPR012939 | Glycosyl hydrolase family 92 | 336 | 715 | 3.5E-103 |
| Pfam | PF07971 | Glycosyl hydrolase family 92 | IPR012939 | Glycosyl hydrolase family 92 | 742 | 858 | 8.9E-27 |
| Gene3D | G3DSA:3.30.2080.10 | 365 | 879 | 2.9E-125 | |||
| TIGRFAM | TIGR01180 | aman2_put: putative alpha-1,2-mannosidase | IPR005887 | Alpha-1,2-mannosidase, putative | 63 | 716 | 5.1E-121 |
| Gene3D | G3DSA:2.70.98.10 | IPR014718 | Glycoside hydrolase-type carbohydrate-binding | 61 | 364 | 5.4E-75 | |
| ProSiteProfiles | PS51257 | Prokaryotic membrane lipoprotein lipid attachment site profile. | 1 | 18 | 6.0 | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 40 | 67 | - | ||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 40 | 56 | - | ||
| SUPERFAMILY | SSF48208 | IPR008928 | Six-hairpin glycosidase superfamily | 335 | 844 | 4.93E-16 | |
| Pfam | PF17678 | Glycosyl hydrolase family 92 N-terminal domain | IPR041371 | Glycosyl hydrolase family 92 N-terminal domain | 70 | 329 | 4.3E-59 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.