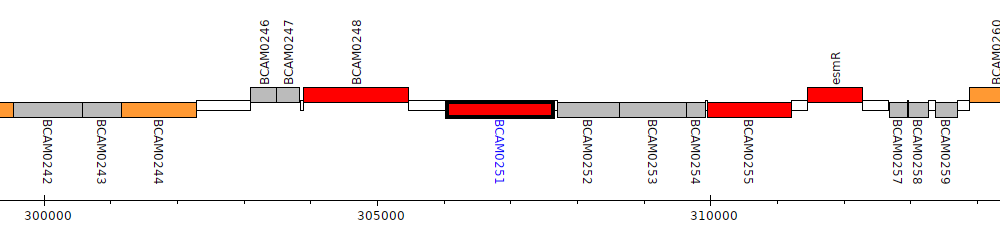
Burkholderia cenocepacia J2315, BCAM0251
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00501
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00154 | AMP-binding signature | IPR020459 | AMP-binding | 164 | 175 | 8.8E-5 |
| CDD | cd05922 | FACL_like_6 | 28 | 515 | 1.22246E-136 | ||
| ProSitePatterns | PS00455 | Putative AMP-binding domain signature. | IPR020845 | AMP-binding, conserved site | 169 | 180 | - |
| Gene3D | G3DSA:3.30.300.30 | 419 | 524 | 1.0E-25 | |||
| SUPERFAMILY | SSF56801 | 3 | 520 | 8.24E-135 | |||
| Gene3D | G3DSA:3.40.50.12780 | IPR042099 | AMP-dependent synthetase-like superfamily | 1 | 418 | 3.7E-99 | |
| TIGRFAM | TIGR03098 | ligase_PEP_1: acyl-CoA ligase (AMP-forming), exosortase A-associated | IPR017529 | Acyl-CoA ligase (AMP-forming), exosortase system type 1 associated | 2 | 515 | 1.2E-210 |
| Pfam | PF13193 | AMP-binding enzyme C-terminal domain | IPR025110 | AMP-binding enzyme, C-terminal domain | 434 | 508 | 1.5E-15 |
| Pfam | PF00501 | AMP-binding enzyme | IPR000873 | AMP-dependent synthetase/ligase | 2 | 425 | 6.8E-88 |
| PRINTS | PR00154 | AMP-binding signature | IPR020459 | AMP-binding | 176 | 184 | 8.8E-5 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.