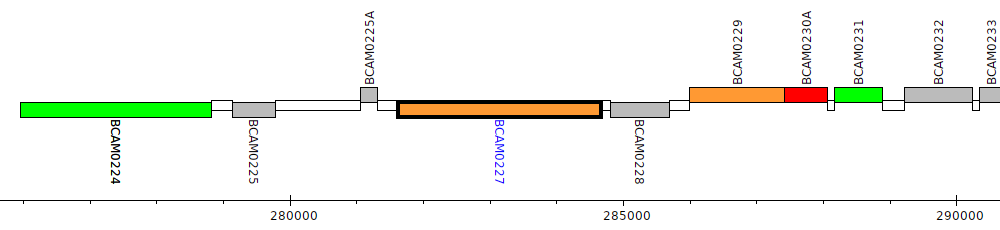
Burkholderia cenocepacia J2315, BCAM0227
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0052100 | intraspecies quorum sensing | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
||
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000160 | phosphorelay signal transduction system |
Inferred from Sequence Model
Term mapped from: InterPro:SSF47226
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000155 | phosphorelay sensor kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00388
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PR00344
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SM00388
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj02020 | Two-component system | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.450.20 | 394 | 518 | 1.9E-5 | |||
| Coils | Coil | 509 | 529 | - | |||
| CDD | cd00082 | HisKA | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 533 | 590 | 2.28554E-15 |
| SUPERFAMILY | SSF52172 | IPR011006 | CheY-like superfamily | 781 | 909 | 1.57E-35 | |
| Gene3D | G3DSA:1.10.287.130 | 519 | 595 | 3.2E-24 | |||
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 717 | 735 | 1.6E-16 |
| Gene3D | G3DSA:3.40.50.2300 | 779 | 913 | 1.8E-35 | |||
| Pfam | PF00072 | Response regulator receiver domain | IPR001789 | Signal transduction response regulator, receiver domain | 783 | 889 | 9.5E-26 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 32 | - | ||
| SMART | SM00448 | IPR001789 | Signal transduction response regulator, receiver domain | 781 | 892 | 1.4E-31 | |
| CDD | cd00156 | REC | IPR001789 | Signal transduction response regulator, receiver domain | 784 | 888 | 4.68411E-31 |
| SUPERFAMILY | SSF55874 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 583 | 758 | 6.29E-46 | |
| Gene3D | G3DSA:3.30.565.10 | IPR036890 | Histidine kinase/HSP90-like ATPase superfamily | 596 | 759 | 2.9E-50 | |
| CDD | cd16922 | HATPase_EvgS-ArcB-TorS-like | 646 | 755 | 1.91461E-53 | ||
| ProSiteProfiles | PS50110 | Response regulatory domain profile. | IPR001789 | Signal transduction response regulator, receiver domain | 782 | 896 | 40.28 |
| SUPERFAMILY | SSF47384 | IPR036097 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain superfamily | 514 | 596 | 2.35E-20 | |
| SMART | SM00388 | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 529 | 594 | 1.1E-21 | |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 700 | 710 | 1.6E-16 |
| ProSiteProfiles | PS50109 | Histidine kinase domain profile. | IPR005467 | Histidine kinase domain | 536 | 757 | 47.824 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 682 | 696 | 1.6E-16 |
| Pfam | PF02518 | Histidine kinase-, DNA gyrase B-, and HSP90-like ATPase | IPR003594 | Histidine kinase/HSP90-like ATPase | 641 | 756 | 7.4E-28 |
| SMART | SM00387 | IPR003594 | Histidine kinase/HSP90-like ATPase | 641 | 757 | 7.2E-37 | |
| Pfam | PF00512 | His Kinase A (phospho-acceptor) domain | IPR003661 | Signal transduction histidine kinase, dimerisation/phosphoacceptor domain | 529 | 594 | 4.4E-19 |
| PRINTS | PR00344 | Bacterial sensor protein C-terminal signature | IPR004358 | Signal transduction histidine kinase-related protein, C-terminal | 741 | 754 | 1.6E-16 |
| SUPERFAMILY | SSF47226 | IPR036641 | HPT domain superfamily | 878 | 991 | 5.76E-8 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.