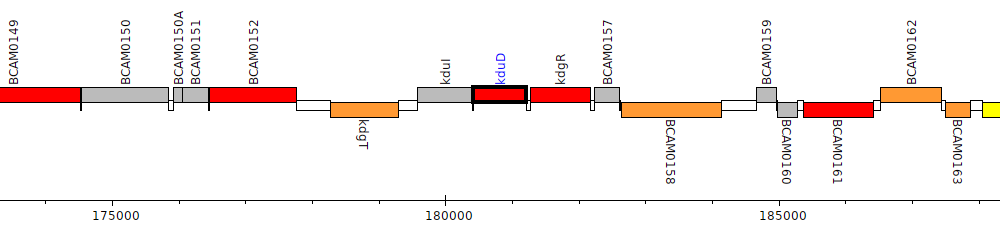
Burkholderia cenocepacia J2315, BCAM0155 (kduD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01832
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008678 | 2-deoxy-D-gluconate 3-dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01832
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01832
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00061
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00040 | Pentose and glucuronate interconversions | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00040 | Pentose and glucuronate interconversions | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | 4-deoxy-L-<i>threo</i>-hex-4-enopyranuronate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 3,6-anhydro-α-L-galactopyranose degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR01832 | kduD: 2-deoxy-D-gluconate 3-dehydrogenase | IPR011286 | 2-dehydro-3-deoxy-D-gluconate 5-dehydrogenase | 21 | 268 | 1.3E-126 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 173 | 192 | 2.9E-44 |
| SUPERFAMILY | SSF51735 | IPR036291 | NAD(P)-binding domain superfamily | 20 | 267 | 1.3E-78 | |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 229 | 249 | 2.9E-44 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 194 | 211 | 2.9E-44 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 99 | 110 | 9.6E-14 |
| ProSitePatterns | PS00061 | Short-chain dehydrogenases/reductases family signature. | IPR020904 | Short-chain dehydrogenase/reductase, conserved site | 160 | 188 | - |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 147 | 163 | 2.9E-44 |
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 99 | 110 | 2.9E-44 |
| Pfam | PF13561 | Enoyl-(Acyl carrier protein) reductase | 33 | 265 | 1.7E-59 | ||
| PRINTS | PR00081 | Glucose/ribitol dehydrogenase family signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 27 | 44 | 2.9E-44 |
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 173 | 192 | 9.6E-14 |
| Gene3D | G3DSA:3.40.50.720 | 17 | 268 | 1.7E-83 | |||
| PRINTS | PR00080 | Short-chain dehydrogenase/reductase (SDR) superfamily signature | IPR002347 | Short-chain dehydrogenase/reductase SDR | 153 | 161 | 9.6E-14 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.