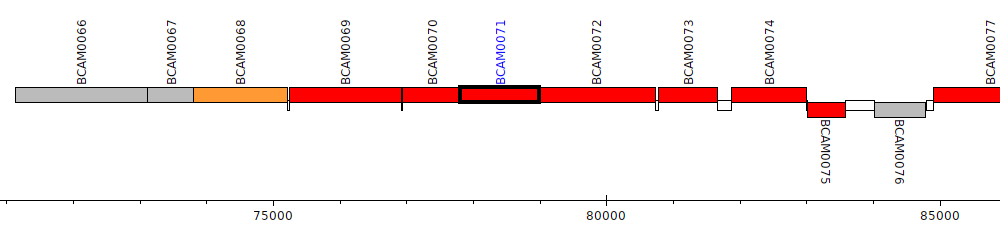
Burkholderia cenocepacia J2315, BCAM0071
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009063 | cellular amino acid catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS00909
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF13378 | Enolase C-terminal domain-like | IPR029065 | Enolase C-terminal domain-like | 157 | 362 | 9.5E-38 |
| SMART | SM00922 | IPR013342 | Mandelate racemase/muconate lactonizing enzyme, C-terminal | 153 | 244 | 2.8E-11 | |
| Gene3D | G3DSA:3.30.390.10 | IPR029017 | Enolase-like, N-terminal | 4 | 120 | 4.1E-70 | |
| Gene3D | G3DSA:3.20.20.120 | IPR036849 | Enolase-like, C-terminal domain superfamily | 121 | 361 | 4.1E-70 | |
| ProSitePatterns | PS00909 | Mandelate racemase / muconate lactonizing enzyme family signature 2. | IPR018110 | Mandelate racemase/muconate lactonizing enzyme, conserved site | 194 | 225 | - |
| SUPERFAMILY | SSF54826 | 3 | 128 | 6.48E-12 | |||
| SFLD | SFLDG00180 | muconate cycloisomerase | 3 | 362 | 0.0 | ||
| SFLD | SFLDS00001 | Enolase | 3 | 362 | 0.0 | ||
| SUPERFAMILY | SSF51604 | IPR036849 | Enolase-like, C-terminal domain superfamily | 160 | 362 | 3.8E-38 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.