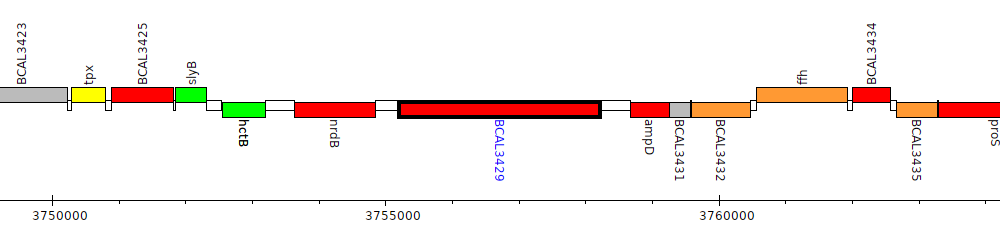
Burkholderia cenocepacia J2315, BCAL3429
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004748 | ribonucleoside-diphosphate reductase activity, thioredoxin disulfide as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:PF00317
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00317
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:PR01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PR01183
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | guanosine deoxyribonucleotides <i>de novo</i> biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | guanosine deoxyribonucleotides <i>de novo</i> biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00983 | Drug metabolism - other enzymes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00240 | Pyrimidine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | pyrimidine deoxyribonucleotides biosynthesis from CTP | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00480 | Glutathione metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | pyrimidine deoxyribonucleotides <i>de novo</i> biosynthesis IV | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyrimidine deoxyribonucleotides <i>de novo</i> biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | pyrimidine deoxyribonucleotides <i>de novo</i> biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adenosine deoxyribonucleotides <i>de novo</i> biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adenosine deoxyribonucleotides <i>de novo</i> biosynthesis II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01183 | Ribonucleotide reductase large chain signature | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 451 | 470 | 6.6E-44 |
| Gene3D | G3DSA:3.20.70.20 | 230 | 926 | 1.1E-245 | |||
| Pfam | PF00317 | Ribonucleotide reductase, all-alpha domain | IPR013509 | Ribonucleotide reductase large subunit, N-terminal | 299 | 372 | 2.8E-21 |
| PRINTS | PR01183 | Ribonucleotide reductase large chain signature | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 666 | 688 | 6.6E-44 |
| Coils | Coil | 1002 | 1003 | - | |||
| ProSitePatterns | PS00089 | Ribonucleotide reductase large subunit signature. | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 759 | 781 | - |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 28 | - | ||
| ProSiteProfiles | PS51161 | ATP-cone domain profile. | IPR005144 | ATP-cone domain | 158 | 247 | 14.937 |
| Pfam | PF02867 | Ribonucleotide reductase, barrel domain | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 376 | 915 | 1.1E-182 |
| ProSiteProfiles | PS51161 | ATP-cone domain profile. | IPR005144 | ATP-cone domain | 40 | 140 | 19.737 |
| PRINTS | PR01183 | Ribonucleotide reductase large chain signature | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 770 | 797 | 6.6E-44 |
| PRINTS | PR01183 | Ribonucleotide reductase large chain signature | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 694 | 717 | 6.6E-44 |
| Pfam | PF03477 | ATP cone domain | IPR005144 | ATP-cone domain | 41 | 137 | 8.2E-19 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1 | 22 | - | ||
| PRINTS | PR01183 | Ribonucleotide reductase large chain signature | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 586 | 597 | 6.6E-44 |
| PRINTS | PR01183 | Ribonucleotide reductase large chain signature | IPR000788 | Ribonucleotide reductase large subunit, C-terminal | 628 | 651 | 6.6E-44 |
| SUPERFAMILY | SSF51998 | 375 | 918 | 3.05E-180 | |||
| SUPERFAMILY | SSF48168 | IPR008926 | Ribonucleotide reductase R1 subunit, N-terminal | 163 | 374 | 2.62E-63 | |
| CDD | cd01679 | RNR_I | 324 | 918 | 0.0 | ||
| TIGRFAM | TIGR02506 | NrdE_NrdA: ribonucleoside-diphosphate reductase, alpha subunit | IPR013346 | Ribonucleotide reductase, class I , alpha subunit | 302 | 918 | 1.3E-210 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.