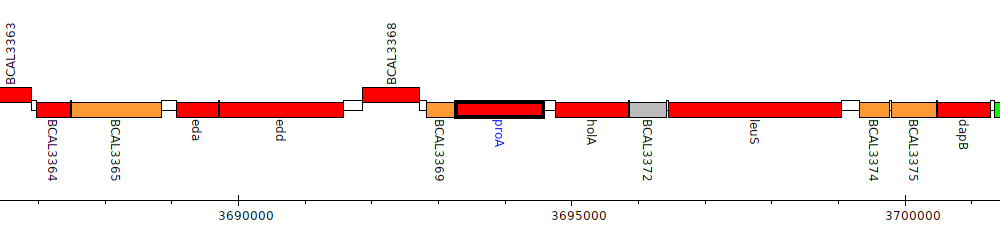
Burkholderia cenocepacia J2315, BCAL3370 (proA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006561 | proline biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00412
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00171
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00171
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050661 | NADP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000151
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.309.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004350 | glutamate-5-semialdehyde dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00412
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00332 | Carbapenem biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00330 | Arginine and proline metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00332 | Carbapenem biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | L-<i>N<sup>δ</sup></i>-acetylornithine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-proline biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00330 | Arginine and proline metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.605.10 | IPR016162 | Aldehyde dehydrogenase, N-terminal | 25 | 431 | 1.2E-169 | |
| Hamap | MF_00412 | Gamma-glutamyl phosphate reductase [proA]. | IPR000965 | GPR domain | 18 | 434 | 41.362 |
| Pfam | PF00171 | Aldehyde dehydrogenase family | IPR015590 | Aldehyde dehydrogenase domain | 24 | 325 | 6.9E-17 |
| ProSitePatterns | PS01223 | Gamma-glutamyl phosphate reductase signature. | IPR020593 | Gamma-glutamyl phosphate reductase GPR, conserved site | 341 | 362 | - |
| CDD | cd07079 | ALDH_F18-19_ProA-GPR | IPR000965 | GPR domain | 20 | 429 | 0.0 |
| Coils | Coil | 42 | 62 | - | |||
| TIGRFAM | TIGR00407 | proA: glutamate-5-semialdehyde dehydrogenase | IPR000965 | GPR domain | 26 | 424 | 3.8E-149 |
| PIRSF | PIRSF000151 | IPR012134 | Glutamate-5-semialdehyde dehydrogenase | 11 | 435 | 6.7E-175 | |
| Pfam | PF00171 | Aldehyde dehydrogenase family | IPR015590 | Aldehyde dehydrogenase domain | 327 | 397 | 4.4E-8 |
| Gene3D | G3DSA:3.40.309.10 | IPR016163 | Aldehyde dehydrogenase, C-terminal | 239 | 392 | 1.2E-169 | |
| SUPERFAMILY | SSF53720 | IPR016161 | Aldehyde/histidinol dehydrogenase | 18 | 432 | 5.89E-118 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.