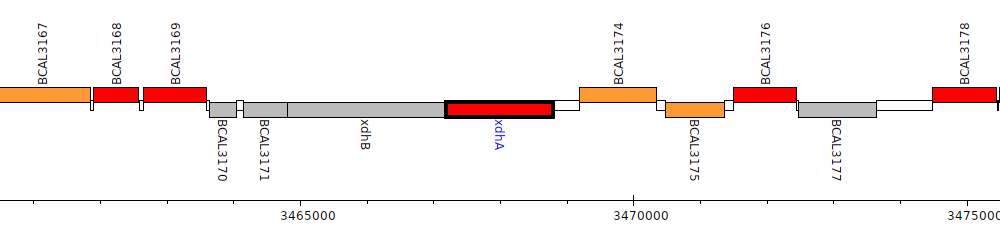
Burkholderia cenocepacia J2315, BCAL3173 (xdhA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004854 | xanthine dehydrogenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02963
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0071949 | FAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS51387
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01799
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051537 | 2 iron, 2 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00197
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02963
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00941
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF54292
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF54292
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004855 | xanthine oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02963
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00941
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | purine nucleobases degradation II (anaerobic) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | theophylline degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | adenosine nucleotides degradation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | caffeine degradation III (bacteria, via demethylation) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | inosine 5'-phosphate degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | guanosine nucleotides degradation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | guanosine nucleotides degradation II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | guanosine nucleotides degradation III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd00207 | fer2 | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 19 | 72 | 6.78454E-4 |
| ProSiteProfiles | PS51387 | PCMH-type FAD-binding domain profile. | IPR016166 | FAD-binding domain, PCMH-type | 220 | 393 | 19.002 |
| SUPERFAMILY | SSF56176 | IPR036318 | FAD-binding, type PCMH-like superfamily | 223 | 390 | 1.42E-52 | |
| SUPERFAMILY | SSF54292 | IPR036010 | 2Fe-2S ferredoxin-like superfamily | 5 | 89 | 8.08E-16 | |
| SMART | SM01092 | IPR005107 | CO dehydrogenase flavoprotein, C-terminal | 401 | 502 | 4.2E-37 | |
| SUPERFAMILY | SSF55447 | IPR036683 | CO dehydrogenase flavoprotein, C-terminal domain superfamily | 399 | 505 | 6.67E-35 | |
| ProSitePatterns | PS00197 | 2Fe-2S ferredoxin-type iron-sulfur binding region signature. | IPR006058 | 2Fe-2S ferredoxin, iron-sulphur binding site | 42 | 50 | - |
| Pfam | PF01799 | [2Fe-2S] binding domain | IPR002888 | [2Fe-2S]-binding | 85 | 171 | 2.5E-31 |
| Gene3D | G3DSA:3.30.390.50 | 401 | 503 | 1.3E-29 | |||
| PIRSF | PIRSF036557 | IPR012175 | Xanthine dehydrogenase, small subunit, bacteria | 2 | 515 | 5.2E-212 | |
| Pfam | PF00941 | FAD binding domain in molybdopterin dehydrogenase | IPR002346 | Molybdopterin dehydrogenase, FAD-binding | 227 | 390 | 7.0E-49 |
| Gene3D | G3DSA:3.10.20.30 | IPR012675 | Beta-grasp domain superfamily | 1 | 88 | 1.7E-25 | |
| Gene3D | G3DSA:1.10.150.120 | 89 | 199 | 1.0E-31 | |||
| Gene3D | G3DSA:3.30.465.10 | IPR016169 | FAD-binding, type PCMH, subdomain 2 | 278 | 390 | 1.2E-31 | |
| Pfam | PF03450 | CO dehydrogenase flavoprotein C-terminal domain | IPR005107 | CO dehydrogenase flavoprotein, C-terminal | 401 | 501 | 6.5E-31 |
| Gene3D | G3DSA:3.30.43.10 | IPR016167 | FAD-binding, type PCMH, subdomain 1 | 228 | 277 | 1.3E-20 | |
| TIGRFAM | TIGR02963 | xanthine_xdhA: xanthine dehydrogenase, small subunit | IPR014307 | Xanthine dehydrogenase, small subunit | 5 | 502 | 3.8E-214 |
| SUPERFAMILY | SSF47741 | IPR036884 | [2Fe-2S]-binding domain superfamily | 97 | 181 | 5.5E-32 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.