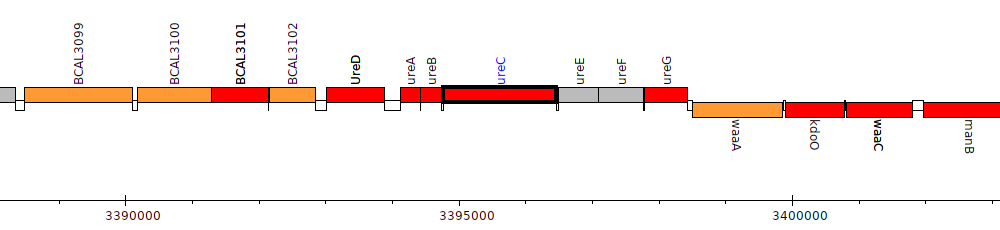
Burkholderia cenocepacia J2315, BCAL3106 (ureC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01979
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009039 | urease activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00145
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006807 | nitrogen compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01953
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016810 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.30.40.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016151 | nickel cation binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01953
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00791 | Atrazine degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00230 | Purine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00330 | Arginine and proline metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00791 | Atrazine degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | urea degradation II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00220 | Arginine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.20.20.140 | 130 | 568 | 0.0 | |||
| ProSitePatterns | PS00145 | Urease active site. | IPR017950 | Urease active site | 318 | 334 | - |
| PRINTS | PR01752 | Urea amidohydrolase (urease) protein signature | IPR005848 | Urease, alpha subunit | 335 | 351 | 1.3E-34 |
| SUPERFAMILY | SSF51338 | IPR011059 | Metal-dependent hydrolase, composite domain superfamily | 4 | 178 | 3.78E-62 | |
| Pfam | PF00449 | Urease alpha-subunit, N-terminal domain | IPR011612 | Urease alpha-subunit, N-terminal domain | 4 | 120 | 4.7E-56 |
| ProSiteProfiles | PS51368 | Urease domain profile. | IPR017951 | Urease alpha subunit, C-terminal | 130 | 568 | 239.15 |
| SUPERFAMILY | SSF51556 | IPR032466 | Metal-dependent hydrolase | 130 | 568 | 4.77E-205 | |
| PRINTS | PR01752 | Urea amidohydrolase (urease) protein signature | IPR005848 | Urease, alpha subunit | 296 | 313 | 1.3E-34 |
| ProSitePatterns | PS01120 | Urease nickel ligands signature. | IPR029754 | Urease nickel binding site | 128 | 141 | - |
| Gene3D | G3DSA:2.30.40.10 | IPR011059 | Metal-dependent hydrolase, composite domain superfamily | 5 | 481 | 0.0 | |
| TIGRFAM | TIGR01792 | urease_alph: urease, alpha subunit | IPR005848 | Urease, alpha subunit | 5 | 568 | 7.0E-301 |
| SUPERFAMILY | SSF51338 | IPR011059 | Metal-dependent hydrolase, composite domain superfamily | 423 | 478 | 4.01E-8 | |
| CDD | cd00375 | Urease_alpha | IPR005848 | Urease, alpha subunit | 4 | 567 | 0.0 |
| PRINTS | PR01752 | Urea amidohydrolase (urease) protein signature | IPR005848 | Urease, alpha subunit | 428 | 441 | 1.3E-34 |
| Pfam | PF01979 | Amidohydrolase family | IPR006680 | Amidohydrolase-related | 126 | 454 | 1.2E-81 |
| Hamap | MF_01953 | Urease subunit alpha [ureC]. | IPR005848 | Urease, alpha subunit | 3 | 568 | 57.897 |
| PRINTS | PR01752 | Urea amidohydrolase (urease) protein signature | IPR005848 | Urease, alpha subunit | 402 | 417 | 1.3E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.