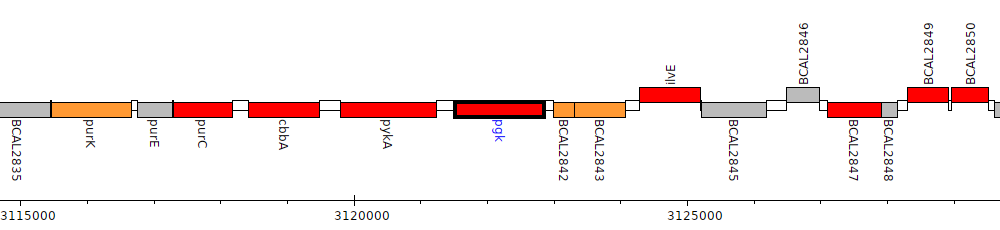
Burkholderia cenocepacia J2315, BCAL2841 (pgk)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006096 | glycolytic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00162
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004618 | phosphoglycerate kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00162
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | glycolysis IV (plant cytosol) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glycolysis II (from fructose 6-phosphate) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | 1-butanol autotrophic biosynthesis (engineered) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00010 | Glycolysis / Gluconeogenesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00010 | Glycolysis / Gluconeogenesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | Entner-Doudoroff pathway I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00710 | Carbon fixation in photosynthetic organisms | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | superpathway of glucose and xylose degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00710 | Carbon fixation in photosynthetic organisms | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glycerol degradation to butanol | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 365 | 390 | 1.2E-79 |
| Hamap | MF_00145 | Phosphoglycerate kinase [pgk]. | IPR001576 | Phosphoglycerate kinase | 59 | 445 | 39.312 |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 243 | 262 | 1.2E-79 |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 398 | 409 | 1.2E-79 |
| SUPERFAMILY | SSF53748 | IPR036043 | Phosphoglycerate kinase superfamily | 65 | 445 | 9.19E-142 | |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 189 | 211 | 1.2E-79 |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 65 | 81 | 1.2E-79 |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 87 | 109 | 1.2E-79 |
| Pfam | PF00162 | Phosphoglycerate kinase | IPR001576 | Phosphoglycerate kinase | 65 | 435 | 4.2E-142 |
| PIRSF | PIRSF000724 | IPR001576 | Phosphoglycerate kinase | 47 | 448 | 9.5E-141 | |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 220 | 242 | 1.2E-79 |
| Gene3D | G3DSA:3.40.50.1260 | IPR015824 | Phosphoglycerate kinase, N-terminal | 222 | 435 | 1.0E-148 | |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 161 | 176 | 1.2E-79 |
| Gene3D | G3DSA:3.40.50.1260 | IPR015824 | Phosphoglycerate kinase, N-terminal | 65 | 443 | 1.0E-148 | |
| PRINTS | PR00477 | Phosphoglycerate kinase family signature | IPR001576 | Phosphoglycerate kinase | 421 | 438 | 1.2E-79 |
| ProSitePatterns | PS00111 | Phosphoglycerate kinase signature. | IPR015911 | Phosphoglycerate kinase, conserved site | 70 | 80 | - |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.