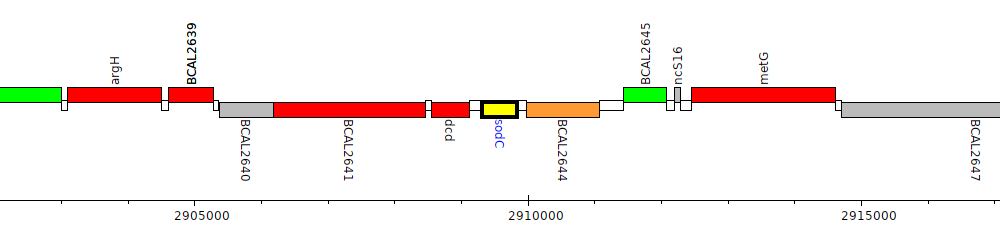
Burkholderia cenocepacia J2315, BCAL2643 (sodC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019430 | removal of superoxide radicals |
Inferred from Sequence or Structural Similarity
Term mapped from: locus_tag:BURCENK562V_RS26530
|
ECO:0000250 sequence similarity evidence used in manual assertion |
17325048 | Reviewed by curator |
| Molecular Function | GO:0004784 | superoxide dismutase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
16997956 | Reviewed by curator |
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.60.40.200
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006801 | superoxide metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.60.40.200
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | ethylene biosynthesis III (microbes) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF49329 | IPR036423 | Superoxide dismutase-like, copper/zinc binding domain superfamily | 44 | 177 | 6.28E-30 | |
| Gene3D | G3DSA:2.60.40.200 | IPR036423 | Superoxide dismutase-like, copper/zinc binding domain superfamily | 29 | 178 | 2.1E-29 | |
| Pfam | PF00080 | Copper/zinc superoxide dismutase (SODC) | IPR001424 | Superoxide dismutase, copper/zinc binding domain | 50 | 177 | 7.1E-25 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.