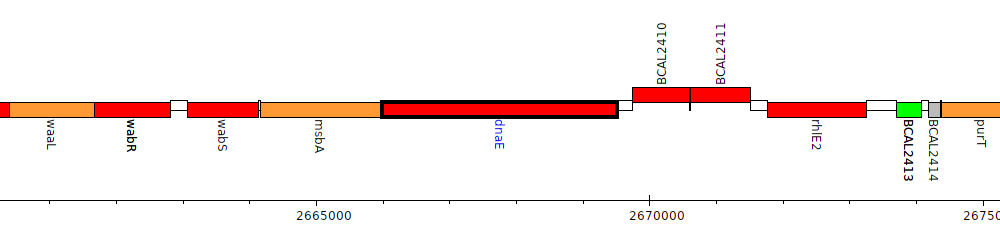
Burkholderia cenocepacia J2315, BCAL2409 (dnaE)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009244 | lipopolysaccharide core region biosynthetic process | Inferred from Genomic Context | ECO:0000317 genomic context evidence used in manual assertion |
19525227 | Reviewed by curator |
| Biological Process | GO:0009244 | lipopolysaccharide core region biosynthetic process | Inferred from Genomic Context | ECO:0000177 genomic context evidence |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF89550
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008408 | 3'-5' exonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00594
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006260 | DNA replication |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00594
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF01336
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03440 | Homologous recombination | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03430 | Mismatch repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03030 | DNA replication | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF17657 | Bacterial DNA polymerase III alpha subunit finger domain | IPR040982 | DNA polymerase III alpha subunit finger domain | 565 | 730 | 2.9E-64 |
| Pfam | PF01336 | OB-fold nucleic acid binding domain | IPR004365 | OB-fold nucleic acid binding domain, AA-tRNA synthetase-type | 988 | 1048 | 2.3E-6 |
| CDD | cd04485 | DnaE_OBF | 987 | 1071 | 1.2002E-21 | ||
| Coils | Coil | 1177 | 1177 | - | |||
| MobiDBLite | mobidb-lite | consensus disorder prediction | 1099 | 1121 | - | ||
| Pfam | PF02811 | PHP domain | IPR004013 | PHP domain | 8 | 173 | 4.4E-43 |
| Pfam | PF07733 | Bacterial DNA polymerase III alpha NTPase domain | IPR011708 | Bacterial DNA polymerase III, alpha subunit, NTPase domain | 293 | 562 | 6.8E-105 |
| Gene3D | G3DSA:1.10.10.1600 | IPR041931 | Bacterial DNA polymerase III alpha subunit, thumb domain | 435 | 514 | 1.3E-27 | |
| Gene3D | G3DSA:3.20.20.140 | 1 | 270 | 4.1E-86 | |||
| CDD | cd07433 | PHP_PolIIIA_DnaE1 | 5 | 282 | 2.16319E-149 | ||
| Pfam | PF14579 | Helix-hairpin-helix motif | IPR029460 | DNA polymerase, helix-hairpin-helix motif | 804 | 898 | 1.6E-28 |
| SMART | SM00481 | IPR003141 | Polymerase/histidinol phosphatase, N-terminal | 7 | 74 | 2.8E-24 | |
| SUPERFAMILY | SSF89550 | IPR016195 | Polymerase/histidinol phosphatase-like | 6 | 264 | 1.57E-11 | |
| Gene3D | G3DSA:2.40.50.140 | 971 | 1069 | 1.9E-9 | |||
| TIGRFAM | TIGR00594 | polc: DNA polymerase III, alpha subunit | IPR004805 | DNA polymerase III, alpha subunit | 6 | 1022 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.