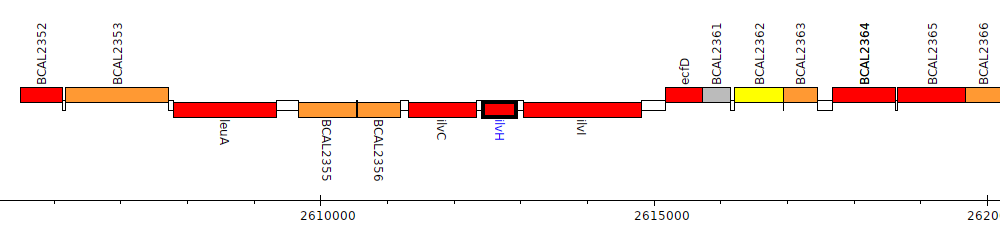
Burkholderia cenocepacia J2315, BCAL2358 (ilvH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:1990610 | acetolactate synthase regulator activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00119
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009082 | branched-chain amino acid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00119
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00290 | Valine, leucine and isoleucine biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00770 | Pantothenate and CoA biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00650 | Butanoate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01210 | 2-Oxocarboxylic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00660 | C5-Branched dibasic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd04878 | ACT_AHAS | IPR039557 | AHAS, ACT domain | 3 | 74 | 1.03554E-31 |
| Gene3D | G3DSA:3.30.70.260 | 1 | 78 | 1.2E-33 | |||
| Gene3D | G3DSA:3.30.70.1150 | IPR027271 | Acetolactate synthase/Transcription factor NikR, C-terminal | 83 | 163 | 5.1E-29 | |
| Pfam | PF13710 | ACT domain | 12 | 73 | 4.8E-15 | ||
| Pfam | PF10369 | Small subunit of acetolactate synthase | IPR019455 | Acetolactate synthase, small subunit, C-terminal | 84 | 157 | 6.0E-26 |
| SUPERFAMILY | SSF55021 | 2 | 76 | 3.23E-21 | |||
| ProSiteProfiles | PS51671 | ACT domain profile. | IPR002912 | ACT domain | 4 | 78 | 14.7 |
| SUPERFAMILY | SSF55021 | 79 | 162 | 7.6E-25 | |||
| TIGRFAM | TIGR00119 | acolac_sm: acetolactate synthase, small subunit | IPR004789 | Acetolactate synthase, small subunit | 1 | 157 | 3.6E-62 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.