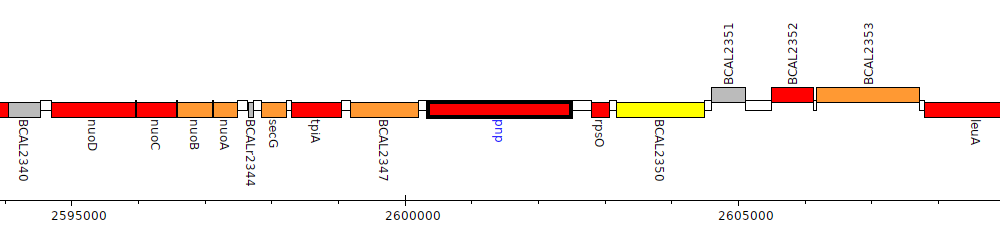
Burkholderia cenocepacia J2315, BCAL2348 (pnp)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006396 | RNA processing |
Inferred from Sequence Model
Term mapped from: InterPro:SSF46915
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004654 | polyribonucleotide nucleotidyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF005499
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS50126
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006402 | mRNA catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF005499
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00013
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj03018 | RNA degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF46915 | IPR036456 | Polyribonucleotide nucleotidyltransferase, RNA-binding domain superfamily | 239 | 326 | 1.39E-16 | |
| Pfam | PF03725 | 3' exoribonuclease family, domain 2 | IPR015847 | Exoribonuclease, phosphorolytic domain 2 | 147 | 210 | 9.7E-12 |
| SMART | SM00316 | IPR022967 | RNA-binding domain, S1 | 625 | 695 | 3.0E-24 | |
| Gene3D | G3DSA:3.30.1370.10 | IPR036612 | K Homology domain, type 1 superfamily | 555 | 626 | 5.2E-21 | |
| SMART | SM00322 | IPR004087 | K Homology domain | 557 | 622 | 7.9E-14 | |
| Pfam | PF03726 | Polyribonucleotide nucleotidyltransferase, RNA binding domain | IPR015848 | Polyribonucleotide nucleotidyltransferase, RNA-binding domain | 242 | 325 | 2.9E-17 |
| Pfam | PF03725 | 3' exoribonuclease family, domain 2 | IPR015847 | Exoribonuclease, phosphorolytic domain 2 | 466 | 534 | 1.2E-4 |
| Pfam | PF01138 | 3' exoribonuclease family, domain 1 | IPR001247 | Exoribonuclease, phosphorolytic domain 1 | 329 | 461 | 3.5E-23 |
| SUPERFAMILY | SSF55666 | IPR036345 | Exoribonuclease, PH domain 2 superfamily | 455 | 553 | 6.03E-28 | |
| Gene3D | G3DSA:3.30.230.70 | IPR027408 | PNPase/RNase PH domain superfamily | 1 | 229 | 2.9E-79 | |
| CDD | cd02393 | PNPase_KH | 558 | 618 | 1.68926E-24 | ||
| PIRSF | PIRSF005499 | IPR012162 | Polyribonucleotide nucleotidyltransferase | 1 | 713 | 3.5E-299 | |
| CDD | cd11363 | RNase_PH_PNPase_1 | 6 | 233 | 3.67067E-125 | ||
| SUPERFAMILY | SSF54791 | IPR036612 | K Homology domain, type 1 superfamily | 559 | 645 | 2.5E-23 | |
| CDD | cd11364 | RNase_PH_PNPase_2 | 328 | 547 | 2.13447E-151 | ||
| SUPERFAMILY | SSF55666 | IPR036345 | Exoribonuclease, PH domain 2 superfamily | 145 | 235 | 2.38E-29 | |
| CDD | cd04472 | S1_PNPase | 627 | 693 | 2.86004E-31 | ||
| Gene3D | G3DSA:3.30.230.70 | IPR027408 | PNPase/RNase PH domain superfamily | 234 | 554 | 1.2E-117 | |
| Gene3D | G3DSA:2.40.50.140 | 627 | 698 | 5.4E-27 | |||
| Pfam | PF00575 | S1 RNA binding domain | IPR003029 | S1 domain | 624 | 695 | 2.4E-17 |
| ProSiteProfiles | PS50084 | Type-1 KH domain profile. | 558 | 617 | 14.445 | ||
| Hamap | MF_01595 | Polyribonucleotide nucleotidyltransferase [pnp]. | IPR012162 | Polyribonucleotide nucleotidyltransferase | 3 | 698 | 30.087 |
| SUPERFAMILY | SSF54211 | IPR020568 | Ribosomal protein S5 domain 2-type fold | 299 | 493 | 2.08E-50 | |
| Pfam | PF01138 | 3' exoribonuclease family, domain 1 | IPR001247 | Exoribonuclease, phosphorolytic domain 1 | 15 | 144 | 4.1E-20 |
| SUPERFAMILY | SSF54211 | IPR020568 | Ribosomal protein S5 domain 2-type fold | 2 | 144 | 1.01E-50 | |
| SUPERFAMILY | SSF50249 | IPR012340 | Nucleic acid-binding, OB-fold | 625 | 704 | 4.93E-21 | |
| ProSiteProfiles | PS50126 | S1 domain profile. | IPR003029 | S1 domain | 627 | 695 | 21.461 |
| Pfam | PF00013 | KH domain | IPR004088 | K Homology domain, type 1 | 561 | 619 | 8.2E-11 |
| TIGRFAM | TIGR03591 | polynuc_phos: polyribonucleotide nucleotidyltransferase | IPR012162 | Polyribonucleotide nucleotidyltransferase | 10 | 696 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.