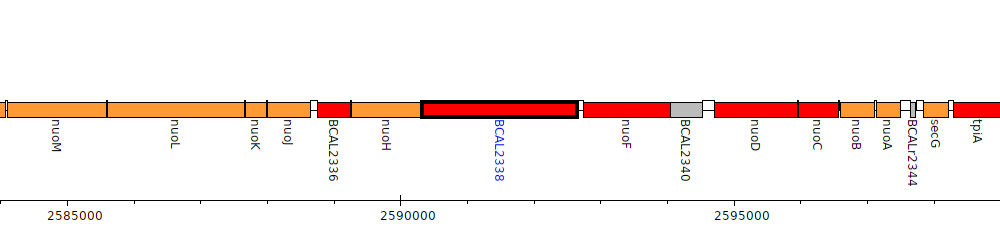
Burkholderia cenocepacia J2315, BCAL2338
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008137 | NADH dehydrogenase (ubiquinone) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00643
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51669
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055114 | oxidation-reduction process |
Inferred from Sequence Model
Term mapped from: InterPro:PS00643
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016651 | oxidoreductase activity, acting on NAD(P)H |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01973
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS51085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0042773 | ATP synthesis coupled electron transport |
Inferred from Sequence Model
Term mapped from: InterPro:PS00643
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS51085
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS00643
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00190 | Oxidative phosphorylation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53706 | 214 | 647 | 1.57E-92 | |||
| SUPERFAMILY | SSF54292 | IPR036010 | 2Fe-2S ferredoxin-like superfamily | 1 | 74 | 4.51E-20 | |
| Pfam | PF13510 | 2Fe-2S iron-sulfur cluster binding domain | 1 | 76 | 8.5E-21 | ||
| ProSitePatterns | PS00643 | Respiratory-chain NADH dehydrogenase 75 Kd subunit signature 3. | IPR000283 | NADH:ubiquinone oxidoreductase, 75kDa subunit, conserved site | 146 | 156 | - |
| SUPERFAMILY | SSF54862 | 75 | 211 | 9.63E-36 | |||
| Pfam | PF00384 | Molybdopterin oxidoreductase | IPR006656 | Molybdopterin oxidoreductase | 273 | 626 | 1.8E-30 |
| ProSitePatterns | PS00641 | Respiratory-chain NADH dehydrogenase 75 Kd subunit signature 1. | IPR000283 | NADH:ubiquinone oxidoreductase, 75kDa subunit, conserved site | 31 | 48 | - |
| ProSiteProfiles | PS51085 | 2Fe-2S ferredoxin-type iron-sulfur binding domain profile. | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 1 | 78 | 9.371 |
| ProSitePatterns | PS00642 | Respiratory-chain NADH dehydrogenase 75 Kd subunit signature 2. | IPR000283 | NADH:ubiquinone oxidoreductase, 75kDa subunit, conserved site | 98 | 110 | - |
| Gene3D | G3DSA:3.10.20.740 | 1 | 123 | 2.3E-42 | |||
| CDD | cd00207 | fer2 | IPR001041 | 2Fe-2S ferredoxin-type iron-sulfur binding domain | 2 | 73 | 1.0133E-9 |
| SMART | SM00929 | IPR019574 | NADH:ubiquinone oxidoreductase, subunit G, iron-sulphur binding | 83 | 123 | 1.0E-19 | |
| ProSiteProfiles | PS51669 | Prokaryotic molybdopterin oxidoreductases 4Fe-4S domain profile. | IPR006963 | Molybdopterin oxidoreductase, 4Fe-4S domain | 216 | 272 | 18.043 |
| Gene3D | G3DSA:3.30.70.20 | 125 | 205 | 2.9E-7 | |||
| Gene3D | G3DSA:3.40.50.740 | 206 | 404 | 2.2E-14 | |||
| Pfam | PF10588 | NADH-ubiquinone oxidoreductase-G iron-sulfur binding region | IPR019574 | NADH:ubiquinone oxidoreductase, subunit G, iron-sulphur binding | 84 | 122 | 3.2E-16 |
| TIGRFAM | TIGR01973 | NuoG: NADH dehydrogenase (quinone), G subunit | IPR010228 | NADH:ubiquinone oxidoreductase, subunit G | 4 | 626 | 4.0E-207 |
| Coils | Coil | 773 | 776 | - | |||
| Gene3D | G3DSA:3.40.50.740 | 420 | 653 | 4.2E-19 | |||
| CDD | cd02772 | MopB_NDH-1_NuoG2 | 220 | 630 | 0.0 | ||
| ProSiteProfiles | PS51839 | His(Cys)3-ligated-type [4Fe-4S] domain profile. | IPR019574 | NADH:ubiquinone oxidoreductase, subunit G, iron-sulphur binding | 78 | 117 | 15.951 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.