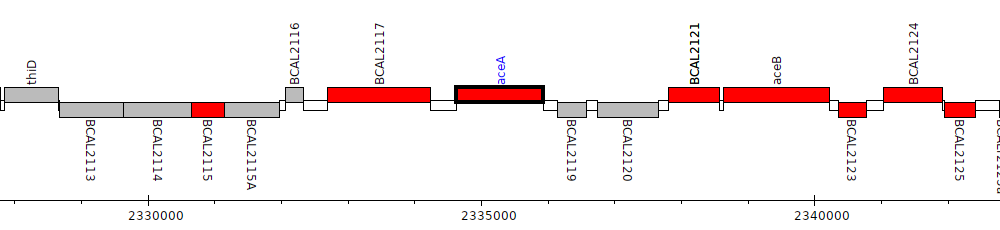
Burkholderia cenocepacia J2315, BCAL2118 (aceA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0019752 | carboxylic acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00463
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF51621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004451 | isocitrate lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00463
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | TCA cycle V (2-oxoglutarate:ferredoxin oxidoreductase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00630 | Glyoxylate and dicarboxylate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00630 | Glyoxylate and dicarboxylate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51621 | IPR015813 | Pyruvate/Phosphoenolpyruvate kinase-like domain superfamily | 4 | 428 | 4.93E-158 | |
| CDD | cd00377 | ICL_PEPM | IPR039556 | ICL/PEPM domain | 53 | 363 | 4.78687E-78 |
| ProSitePatterns | PS00161 | Isocitrate lyase signature. | IPR018523 | Isocitrate lyase/phosphorylmutase, conserved site | 190 | 195 | - |
| TIGRFAM | TIGR01346 | isocit_lyase: isocitrate lyase | IPR006254 | Isocitrate lyase | 7 | 252 | 4.3E-112 |
| Pfam | PF00463 | Isocitrate lyase family | IPR006254 | Isocitrate lyase | 12 | 252 | 6.8E-60 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | 415 | 435 | - | ||
| PIRSF | PIRSF001362 | IPR006254 | Isocitrate lyase | 1 | 431 | 8.3E-239 | |
| Gene3D | G3DSA:3.20.20.60 | IPR040442 | Pyruvate kinase-like domain superfamily | 1 | 412 | 3.2E-173 | |
| Pfam | PF00463 | Isocitrate lyase family | IPR006254 | Isocitrate lyase | 253 | 425 | 3.0E-44 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.