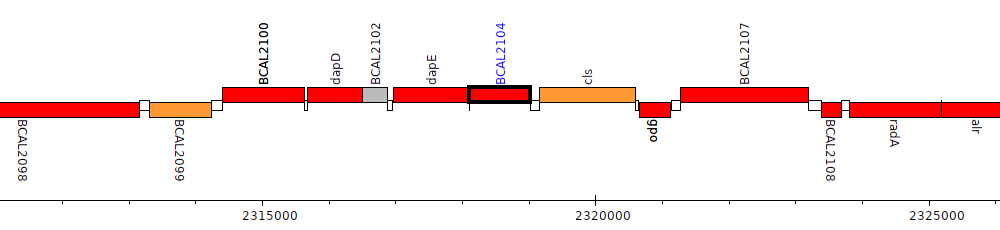
Burkholderia cenocepacia J2315, BCAL2104
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0032259 | methylation |
Inferred from Sequence Model
Term mapped from: InterPro:PS00092
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0032775 | DNA methylation on adenine |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03533
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008276 | protein methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00536
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008168 | methyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF05175
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006479 | protein methylation |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00536
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009007 | site-specific DNA-methyltransferase (adenine-specific) activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03533
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00092
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.150 | 98 | 295 | 6.4E-45 | |||
| Pfam | PF05175 | Methyltransferase small domain | IPR007848 | Methyltransferase small domain | 126 | 213 | 1.5E-11 |
| Gene3D | G3DSA:1.10.8.10 | 7 | 97 | 1.6E-6 | |||
| SUPERFAMILY | SSF53335 | IPR029063 | S-adenosyl-L-methionine-dependent methyltransferase | 8 | 267 | 2.94E-47 | |
| Hamap | MF_02125 | 50S ribosomal protein L3 glutamine methyltransferase [prmB]. | IPR017127 | Ribosomal protein L3-specific, glutamine-N5-methyltransferase | 1 | 300 | 50.804 |
| PIRSF | PIRSF037167 | IPR017127 | Ribosomal protein L3-specific, glutamine-N5-methyltransferase | 1 | 302 | 6.1E-126 | |
| TIGRFAM | TIGR03533 | L3_gln_methyl: protein-(glutamine-N5) methyltransferase, ribosomal protein L3-specific | IPR017127 | Ribosomal protein L3-specific, glutamine-N5-methyltransferase | 6 | 294 | 1.4E-120 |
| ProSitePatterns | PS00092 | N-6 Adenine-specific DNA methylases signature. | IPR002052 | DNA methylase, N-6 adenine-specific, conserved site | 204 | 210 | - |
| TIGRFAM | TIGR00536 | hemK_fam: methyltransferase, HemK family | IPR004556 | Methyltransferase HemK-like | 7 | 270 | 7.5E-64 |
| CDD | cd02440 | AdoMet_MTases | 130 | 262 | 4.65586E-11 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.