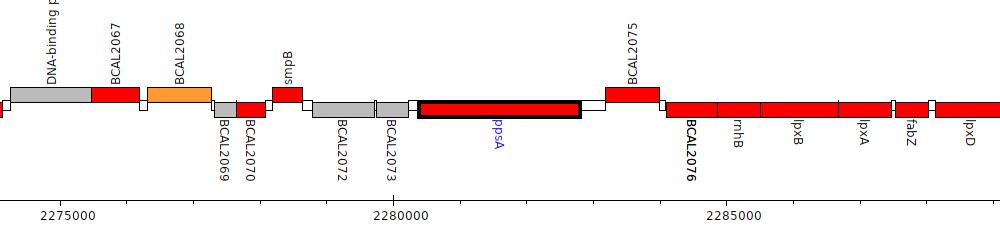
Burkholderia cenocepacia J2315, BCAL2074 (ppsA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1490.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016310 | phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:PF02896
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF51621
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016301 | kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01326
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008986 | pyruvate, water dikinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000854
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016772 | transferase activity, transferring phosphorus-containing groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF02896
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006090 | pyruvate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000854
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | gluconeogenesis II (<i>Methanobacterium thermoautotrophicum</i>) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj00680 | Methane metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00620 | Pyruvate metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | glycolysis II (from fructose 6-phosphate) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00720 | Carbon fixation pathways in prokaryotes | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00680 | Methane metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00620 | Pyruvate metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR01418 | PEP_synth: phosphoenolpyruvate synthase | IPR006319 | Phosphoenolpyruvate synthase | 14 | 798 | 0.0 |
| Gene3D | G3DSA:3.20.20.60 | IPR040442 | Pyruvate kinase-like domain superfamily | 486 | 799 | 5.2E-99 | |
| SUPERFAMILY | SSF56059 | 11 | 393 | 1.53E-108 | |||
| PIRSF | PIRSF000854 | IPR006319 | Phosphoenolpyruvate synthase | 8 | 800 | 0.0 | |
| ProSitePatterns | PS00742 | PEP-utilizing enzymes signature 2. | IPR023151 | PEP-utilising enzyme, conserved site | 702 | 720 | - |
| SUPERFAMILY | SSF52009 | IPR036637 | Phosphohistidine domain superfamily | 356 | 475 | 7.33E-34 | |
| Pfam | PF02896 | PEP-utilising enzyme, TIM barrel domain | IPR000121 | PEP-utilising enzyme, C-terminal | 489 | 796 | 3.3E-60 |
| Pfam | PF00391 | PEP-utilising enzyme, mobile domain | IPR008279 | PEP-utilising enzyme, mobile domain | 392 | 463 | 7.6E-26 |
| Pfam | PF01326 | Pyruvate phosphate dikinase, PEP/pyruvate binding domain | IPR002192 | Pyruvate phosphate dikinase, PEP/pyruvate-binding | 26 | 353 | 5.3E-117 |
| Gene3D | G3DSA:3.30.1490.20 | IPR013815 | ATP-grasp fold, subdomain 1 | 9 | 202 | 9.7E-73 | |
| ProSitePatterns | PS00370 | PEP-utilizing enzymes phosphorylation site signature. | IPR018274 | PEP-utilising enzyme, active site | 422 | 433 | - |
| Gene3D | G3DSA:3.30.470.20 | 203 | 364 | 7.7E-57 | |||
| SUPERFAMILY | SSF51621 | IPR015813 | Pyruvate/Phosphoenolpyruvate kinase-like domain superfamily | 472 | 797 | 1.88E-93 | |
| Gene3D | G3DSA:3.50.30.10 | 365 | 475 | 9.0E-40 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.