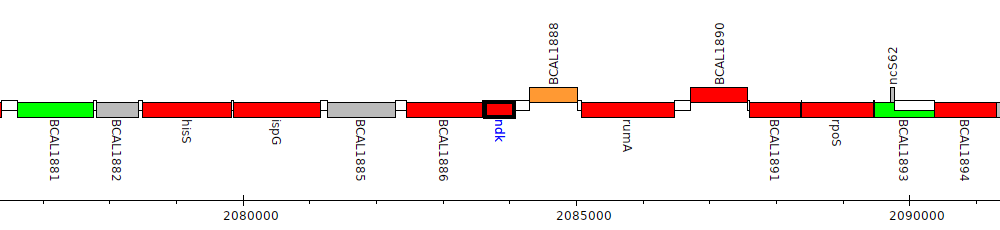
Burkholderia cenocepacia J2315, BCAL1887 (ndk)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004550 | nucleoside diphosphate kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00451
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006228 | UTP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00451
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006165 | nucleoside diphosphate phosphorylation |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00451
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006241 | CTP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00451
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006183 | GTP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00451
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00240 | Pyrimidine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00230 | Purine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01243 | Nucleoside diphosphate kinase signature | IPR001564 | Nucleoside diphosphate kinase | 91 | 107 | 2.9E-37 |
| PRINTS | PR01243 | Nucleoside diphosphate kinase signature | IPR001564 | Nucleoside diphosphate kinase | 6 | 28 | 2.9E-37 |
| PRINTS | PR01243 | Nucleoside diphosphate kinase signature | IPR001564 | Nucleoside diphosphate kinase | 114 | 133 | 2.9E-37 |
| CDD | cd04413 | NDPk_I | 4 | 133 | 2.29496E-75 | ||
| SUPERFAMILY | SSF54919 | IPR036850 | Nucleoside diphosphate kinase-like domain superfamily | 1 | 139 | 5.76E-56 | |
| Gene3D | G3DSA:3.30.70.141 | IPR036850 | Nucleoside diphosphate kinase-like domain superfamily | 1 | 141 | 1.9E-58 | |
| SMART | SM00562 | IPR034907 | Nucleoside diphosphate kinase-like domain | 3 | 140 | 3.0E-78 | |
| PRINTS | PR01243 | Nucleoside diphosphate kinase signature | IPR001564 | Nucleoside diphosphate kinase | 50 | 69 | 2.9E-37 |
| PRINTS | PR01243 | Nucleoside diphosphate kinase signature | IPR001564 | Nucleoside diphosphate kinase | 70 | 87 | 2.9E-37 |
| Pfam | PF00334 | Nucleoside diphosphate kinase | IPR034907 | Nucleoside diphosphate kinase-like domain | 4 | 137 | 2.1E-53 |
| Hamap | MF_00451 | Nucleoside diphosphate kinase [ndk]. | IPR001564 | Nucleoside diphosphate kinase | 3 | 138 | 33.786 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.