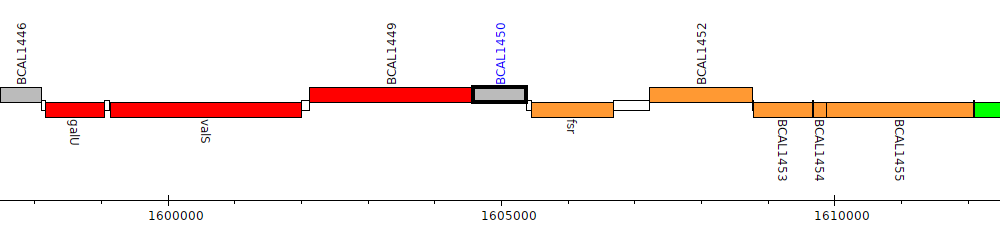
Burkholderia cenocepacia J2315, BCAL1450
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008782 | adenosylhomocysteine nucleosidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01704
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019509 | L-methionine salvage from methylthioadenosine |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01704
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53167
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009164 | nucleoside catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01704
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009116 | nucleoside metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53167
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008930 | methylthioadenosine nucleosidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01704
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00270 | Cysteine and methionine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | <i>S</i>-adenosyl-L-methionine cycle I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | autoinducer AI-2 biosynthesis I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00270 | Cysteine and methionine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | autoinducer AI-2 biosynthesis II (<i>Vibrio</i>) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR01704 | MTA/SAH-Nsdase: MTA/SAH nucleosidase | IPR010049 | MTA/SAH nucleosidase | 16 | 252 | 3.3E-33 |
| SUPERFAMILY | SSF53167 | IPR035994 | Nucleoside phosphorylase superfamily | 15 | 253 | 1.57E-54 | |
| Gene3D | G3DSA:3.40.50.1580 | IPR035994 | Nucleoside phosphorylase superfamily | 11 | 252 | 3.9E-54 | |
| CDD | cd09008 | MTAN | 15 | 256 | 1.02548E-66 | ||
| Pfam | PF01048 | Phosphorylase superfamily | IPR000845 | Nucleoside phosphorylase domain | 15 | 257 | 2.4E-45 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.