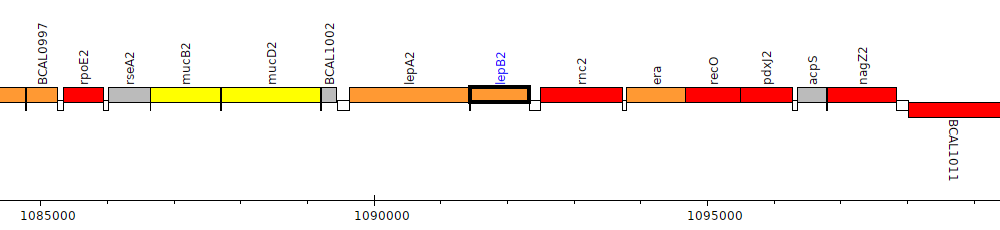
Burkholderia cenocepacia J2315, BCAL1004 (lepB2)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02227
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006508 | proteolysis |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02227
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008236 | serine-type peptidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PS00501
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PS00501
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03060 | Protein export | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd06530 | S26_SPase_I | 96 | 280 | 2.87504E-26 | ||
| ProSitePatterns | PS00761 | Signal peptidases I signature 3. | IPR019758 | Peptidase S26A, signal peptidase I, conserved site | 252 | 265 | - |
| ProSitePatterns | PS00760 | Signal peptidases I lysine active site. | IPR019757 | Peptidase S26A, signal peptidase I, lysine active site | 159 | 171 | - |
| PRINTS | PR00727 | Bacterial leader peptidase 1 (S26A) family signature | IPR000223 | Peptidase S26A, signal peptidase I | 157 | 169 | 7.1E-26 |
| ProSitePatterns | PS00501 | Signal peptidases I serine active site. | IPR019756 | Peptidase S26A, signal peptidase I, serine active site | 102 | 109 | - |
| PRINTS | PR00727 | Bacterial leader peptidase 1 (S26A) family signature | IPR000223 | Peptidase S26A, signal peptidase I | 93 | 109 | 7.1E-26 |
| Pfam | PF00717 | Peptidase S24-like | IPR015927 | Peptidase S24/S26A/S26B/S26C | 99 | 181 | 8.3E-22 |
| Pfam | PF10502 | Signal peptidase, peptidase S26 | IPR019533 | Peptidase S26 | 246 | 284 | 1.6E-6 |
| Gene3D | G3DSA:2.10.109.10 | 91 | 287 | 1.1E-75 | |||
| PRINTS | PR00727 | Bacterial leader peptidase 1 (S26A) family signature | IPR000223 | Peptidase S26A, signal peptidase I | 247 | 266 | 7.1E-26 |
| TIGRFAM | TIGR02227 | sigpep_I_bact: signal peptidase I | IPR000223 | Peptidase S26A, signal peptidase I | 78 | 287 | 1.8E-53 |
| SUPERFAMILY | SSF51306 | IPR036286 | LexA/Signal peptidase-like superfamily | 91 | 293 | 2.62E-72 | |
| Gene3D | G3DSA:2.170.230.10 | 168 | 246 | 1.1E-75 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.