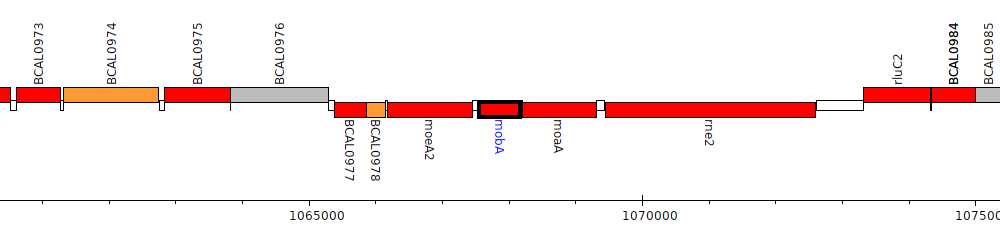
Burkholderia cenocepacia J2315, BCAL0980 (mobA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006777 | Mo-molybdopterin cofactor biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd02503
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd02503
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG (InterPro) | 00790 | Folate biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| MetaCyc | bis(guanylyl molybdenum cofactor) biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | guanylyl molybdenum cofactor biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR02665 | molyb_mobA: molybdenum cofactor guanylyltransferase | IPR013482 | Molybdenum cofactor guanylyltransferase | 9 | 200 | 1.0E-65 |
| SUPERFAMILY | SSF53448 | IPR029044 | Nucleotide-diphospho-sugar transferases | 7 | 199 | 6.24E-31 | |
| Pfam | PF12804 | MobA-like NTP transferase domain | IPR025877 | MobA-like NTP transferase | 11 | 178 | 1.8E-31 |
| CDD | cd02503 | MobA | IPR013482 | Molybdenum cofactor guanylyltransferase | 9 | 199 | 5.27861E-55 |
| Hamap | MF_00316 | Molybdenum cofactor guanylyltransferase [mobA]. | IPR013482 | Molybdenum cofactor guanylyltransferase | 7 | 202 | 27.024 |
| Gene3D | G3DSA:3.90.550.10 | IPR029044 | Nucleotide-diphospho-sugar transferases | 6 | 205 | 8.0E-50 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.