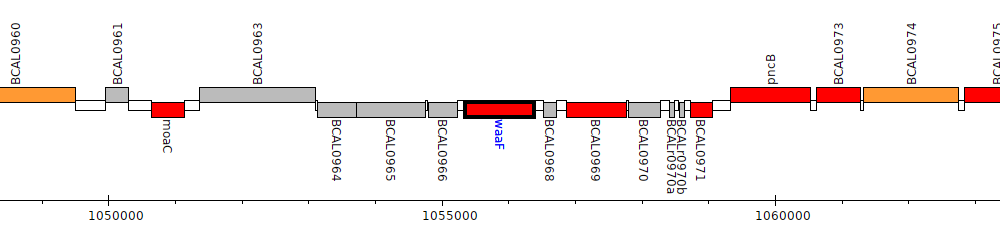
Burkholderia cenocepacia J2315, BCAL0967 (waaF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009244 | lipopolysaccharide core region biosynthetic process | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
19525227 | Reviewed by curator |
| Biological Process | GO:0009103 | lipopolysaccharide biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02195
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016757 | transferase activity, transferring glycosyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PF01075
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00540 | Lipopolysaccharide biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF53756 | 2 | 341 | 3.03E-98 | |||
| CDD | cd03789 | GT9_LPS_heptosyltransferase | 3 | 338 | 1.97338E-79 | ||
| TIGRFAM | TIGR02195 | heptsyl_trn_II: lipopolysaccharide heptosyltransferase II | IPR011910 | ADP-heptose--LPS heptosyltransferase 2 | 3 | 341 | 2.9E-148 |
| Pfam | PF01075 | Glycosyltransferase family 9 (heptosyltransferase) | IPR002201 | Glycosyl transferase, family 9 | 70 | 322 | 1.7E-65 |
| Gene3D | G3DSA:3.40.50.2000 | 3 | 152 | 1.9E-50 | |||
| Gene3D | G3DSA:3.40.50.2000 | 166 | 344 | 3.1E-58 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.