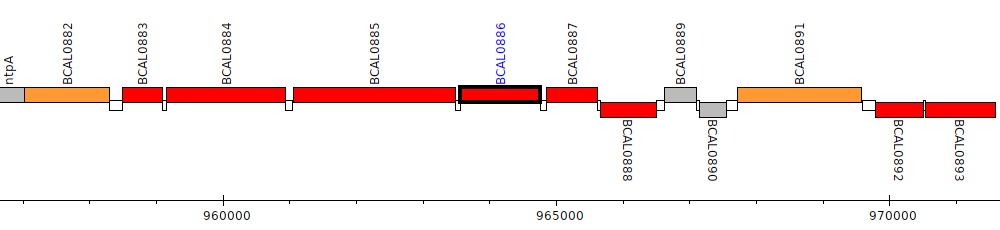
Burkholderia cenocepacia J2315, BCAL0886
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF53901
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016747 | transferase activity, transferring acyl groups other than amino-acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000429
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00071 | Fatty acid degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01212 | Fatty acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00362 | Benzoate degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00281 | Geraniol degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01120 | Microbial metabolism in diverse environments | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00280 | Valine, leucine and isoleucine degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00592 | alpha-Linolenic acid metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd00751 | thiolase | IPR002155 | Thiolase | 9 | 398 | 0.0 |
| SUPERFAMILY | SSF53901 | IPR016039 | Thiolase-like | 277 | 398 | 7.36E-44 | |
| SUPERFAMILY | SSF53901 | IPR016039 | Thiolase-like | 5 | 273 | 4.46E-72 | |
| PIRSF | PIRSF000429 | IPR002155 | Thiolase | 2 | 399 | 3.2E-128 | |
| ProSitePatterns | PS00098 | Thiolases acyl-enzyme intermediate signature. | IPR020615 | Thiolase, acyl-enzyme intermediate active site | 91 | 109 | - |
| ProSitePatterns | PS00737 | Thiolases signature 2. | IPR020613 | Thiolase, conserved site | 345 | 361 | - |
| TIGRFAM | TIGR01930 | AcCoA-C-Actrans: acetyl-CoA C-acyltransferase | IPR002155 | Thiolase | 10 | 397 | 1.8E-138 |
| ProSitePatterns | PS00099 | Thiolases active site. | IPR020610 | Thiolase, active site | 380 | 393 | - |
| Gene3D | G3DSA:3.40.47.10 | IPR016039 | Thiolase-like | 9 | 282 | 6.5E-142 | |
| Pfam | PF02803 | Thiolase, C-terminal domain | IPR020617 | Thiolase, C-terminal | 276 | 398 | 1.5E-44 |
| Gene3D | G3DSA:3.40.47.10 | IPR016039 | Thiolase-like | 133 | 393 | 6.5E-142 | |
| Pfam | PF00108 | Thiolase, N-terminal domain | IPR020616 | Thiolase, N-terminal | 9 | 268 | 1.7E-69 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.