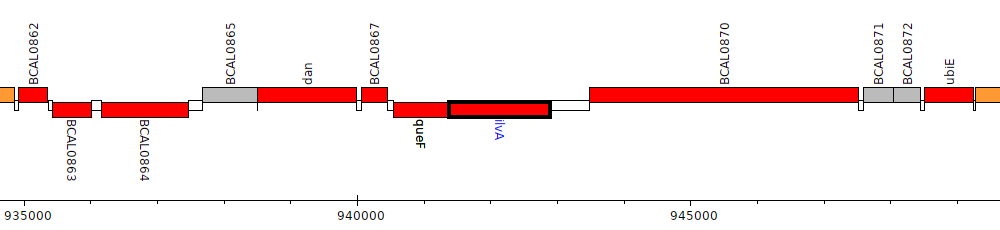
Burkholderia cenocepacia J2315, BCAL0869 (ilvA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004794 | L-threonine ammonia-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01124
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030170 | pyridoxal phosphate binding |
Inferred from Sequence Model
Term mapped from: InterPro:PS00165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009097 | isoleucine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01124
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006520 | cellular amino acid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PS00165
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj00290 | Valine, leucine and isoleucine biosynthesis | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01230 | Biosynthesis of amino acids | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01200 | Carbon metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00260 | Glycine, serine and threonine metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00260 | Glycine, serine and threonine metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00290 | Valine, leucine and isoleucine biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01130 | Biosynthesis of antibiotics | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00291 | Pyridoxal-phosphate dependent enzyme | IPR001926 | Pyridoxal-phosphate dependent enzyme | 18 | 310 | 3.4E-79 |
| CDD | cd01562 | Thr-dehyd | 7 | 314 | 1.01922E-152 | ||
| SUPERFAMILY | SSF53686 | IPR036052 | Tryptophan synthase beta subunit-like PLP-dependent enzyme | 5 | 360 | 6.42E-98 | |
| ProSitePatterns | PS00165 | Serine/threonine dehydratases pyridoxal-phosphate attachment site. | IPR000634 | Serine/threonine dehydratase, pyridoxal-phosphate-binding site | 43 | 56 | - |
| CDD | cd04906 | ACT_ThrD-I_1 | 332 | 415 | 4.87248E-44 | ||
| TIGRFAM | TIGR01124 | ilvA_2Cterm: threonine ammonia-lyase, biosynthetic | IPR005787 | Threonine dehydratase, biosynthetic | 5 | 506 | 2.0E-250 |
| ProSiteProfiles | PS51672 | ACT-like domain profile. | IPR001721 | Threonine dehydratase, ACT-like domain | 427 | 498 | 17.613 |
| Gene3D | G3DSA:3.40.1020.10 | IPR038110 | Threonine dehydratase, ACT-like domain superfamily | 332 | 507 | 7.5E-75 | |
| CDD | cd04907 | ACT_ThrD-I_2 | 426 | 506 | 1.78629E-45 | ||
| Pfam | PF00585 | C-terminal regulatory domain of Threonine dehydratase | IPR001721 | Threonine dehydratase, ACT-like domain | 323 | 412 | 7.7E-31 |
| Gene3D | G3DSA:3.40.50.1100 | 51 | 150 | 1.9E-136 | |||
| Pfam | PF00585 | C-terminal regulatory domain of Threonine dehydratase | IPR001721 | Threonine dehydratase, ACT-like domain | 424 | 506 | 5.8E-24 |
| SUPERFAMILY | SSF55021 | 413 | 506 | 5.49E-27 | |||
| ProSiteProfiles | PS51672 | ACT-like domain profile. | IPR001721 | Threonine dehydratase, ACT-like domain | 333 | 404 | 14.101 |
| Gene3D | G3DSA:3.40.50.1100 | 15 | 321 | 1.9E-136 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.