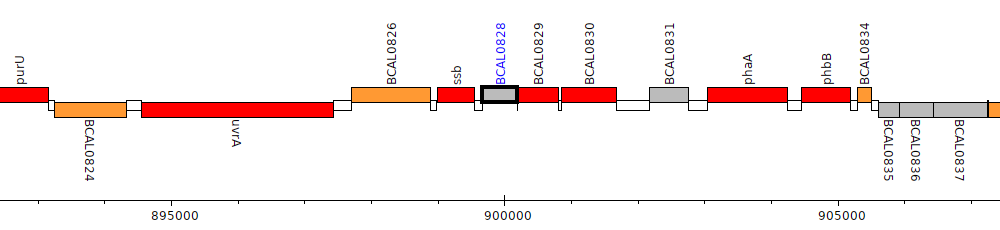
Burkholderia cenocepacia J2315, BCAL0828
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0019104 | DNA N-glycosylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd10028
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006281 | DNA repair |
Inferred from Sequence Model
Term mapped from: InterPro:cd10028
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj03410 | Base excision repair | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52141 | IPR036895 | Uracil-DNA glycosylase-like domain superfamily | 6 | 163 | 1.21E-42 | |
| Gene3D | G3DSA:3.40.470.10 | IPR036895 | Uracil-DNA glycosylase-like domain superfamily | 4 | 172 | 2.3E-62 | |
| SMART | SM00987 | 6 | 163 | 1.9E-4 | |||
| Pfam | PF03167 | Uracil DNA glycosylase superfamily | IPR005122 | Uracil-DNA glycosylase-like | 9 | 161 | 5.9E-25 |
| CDD | cd10028 | UDG_F2_MUG | IPR015637 | G/U mismatch-specific DNA glycosylase | 5 | 164 | 1.52652E-78 |
| SMART | SM00986 | IPR005122 | Uracil-DNA glycosylase-like | 6 | 163 | 1.9E-4 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.