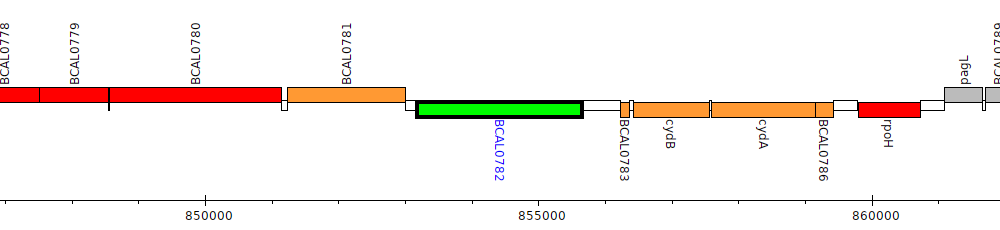
Burkholderia cenocepacia J2315, BCAL0782
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PR00738
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.60.40.290
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004563 | beta-N-acetylhexosaminidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00738
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030247 | polysaccharide binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:2.60.40.290
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0030246 | carbohydrate binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF49384
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | chitin degradation II (Vibrio) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00531 | Glycosaminoglycan degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00520 | Amino sugar and nucleotide sugar metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG (InterPro) | 00511 | Other glycan degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00604 | Glycosphingolipid biosynthesis - ganglio series | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00520 | Amino sugar and nucleotide sugar metabolism | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj01110 | Biosynthesis of secondary metabolites | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj01100 | Metabolic pathways | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | anhydromuropeptides recycling II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | chitin degradation III (Serratia) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG (InterPro) | 00513 | Various types of N-glycan biosynthesis | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG (InterPro) | 00603 | Glycosphingolipid biosynthesis - globo and isoglobo series | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
|
| KEGG | bcj00511 | Other glycan degradation | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 748 | 765 | 5.4E-42 |
| Gene3D | G3DSA:3.30.379.10 | IPR029018 | Beta-hexosaminidase-like, domain 2 | 228 | 334 | 2.4E-14 | |
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 436 | 453 | 5.4E-42 |
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 298 | 318 | 5.4E-42 |
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 530 | 543 | 5.4E-42 |
| Pfam | PF03173 | Putative carbohydrate binding domain | IPR004866 | Chitobiase/beta-hexosaminidases, N-terminal domain | 56 | 203 | 4.5E-33 |
| SUPERFAMILY | SSF81296 | IPR014756 | Immunoglobulin E-set | 783 | 822 | 5.32E-8 | |
| Gene3D | G3DSA:2.60.40.290 | IPR012291 | CBM2, carbohydrate-binding domain superfamily | 47 | 188 | 2.3E-34 | |
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 730 | 746 | 5.4E-42 |
| SUPERFAMILY | SSF51445 | IPR017853 | Glycoside hydrolase superfamily | 337 | 778 | 2.19E-130 | |
| SUPERFAMILY | SSF55545 | IPR029018 | Beta-hexosaminidase-like, domain 2 | 218 | 336 | 9.42E-20 | |
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 331 | 348 | 5.4E-42 |
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 503 | 521 | 5.4E-42 |
| Pfam | PF02838 | Glycosyl hydrolase family 20, domain 2 | IPR015882 | Beta-hexosaminidase, bacterial type, N-terminal | 238 | 334 | 4.2E-12 |
| Pfam | PF00728 | Glycosyl hydrolase family 20, catalytic domain | IPR015883 | Glycoside hydrolase family 20, catalytic domain | 337 | 766 | 2.6E-97 |
| SMART | SM01081 | IPR004866 | Chitobiase/beta-hexosaminidases, N-terminal domain | 54 | 217 | 9.3E-63 | |
| PRINTS | PR00738 | Glycosyl hydrolase family 20 signature | IPR025705 | Beta-hexosaminidase | 360 | 381 | 5.4E-42 |
| Gene3D | G3DSA:3.20.20.80 | 335 | 820 | 5.3E-148 | |||
| SUPERFAMILY | SSF49384 | IPR008965 | CBM2/CBM3, carbohydrate-binding domain superfamily | 52 | 212 | 1.03E-45 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.