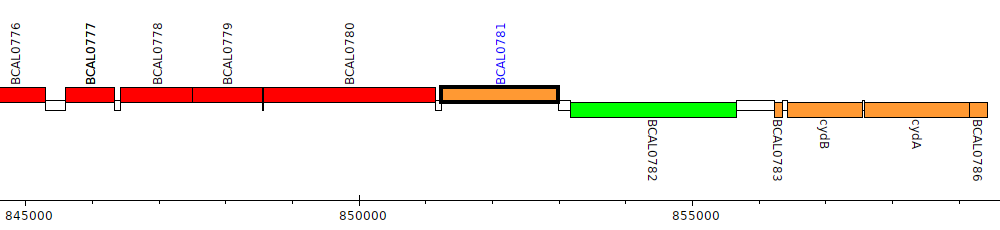
Burkholderia cenocepacia J2315, BCAL0781
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009401 | phosphoenolpyruvate-dependent sugar phosphotransferase system |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55604
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF02378
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016021 | integral component of membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01998
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015572 | N-acetylglucosamine transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01998
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0019866 | organelle inner membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01998
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008982 | protein-N(PI)-phosphohistidine-sugar phosphotransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55604
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | bcj02060 | Phosphotransferase system (PTS) | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | bcj00520 | Amino sugar and nucleotide sugar metabolism | 73.0+/03-31, Mar 15 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| TIGRFAM | TIGR00826 | EIIB_glc: PTS system, glucose-like IIB component | IPR001996 | Phosphotransferase system, IIB component, type 1 | 360 | 461 | 4.3E-16 |
| Pfam | PF00367 | phosphotransferase system, EIIB | IPR018113 | Phosphotransferase system EIIB, cysteine phosphorylation site | 508 | 540 | 3.6E-8 |
| CDD | cd00212 | PTS_IIB_glc | IPR018113 | Phosphotransferase system EIIB, cysteine phosphorylation site | 403 | 472 | 4.40698E-23 |
| ProSitePatterns | PS01035 | PTS EIIB domains cysteine phosphorylation site signature. | IPR018113 | Phosphotransferase system EIIB, cysteine phosphorylation site | 414 | 431 | - |
| Pfam | PF02378 | Phosphotransferase system, EIIC | IPR003352 | Phosphotransferase system, EIIC | 14 | 303 | 3.2E-78 |
| ProSiteProfiles | PS51103 | PTS_EIIC type-1 domain profile. | IPR013013 | Phosphotransferase system, EIIC component, type 1 | 3 | 371 | 62.509 |
| Gene3D | G3DSA:3.30.1360.60 | IPR036878 | Glucose permease domain IIB | 504 | 576 | 1.3E-8 | |
| SUPERFAMILY | SSF55604 | IPR036878 | Glucose permease domain IIB | 507 | 573 | 3.14E-9 | |
| Gene3D | G3DSA:3.30.1360.60 | IPR036878 | Glucose permease domain IIB | 397 | 476 | 1.4E-23 | |
| Pfam | PF00367 | phosphotransferase system, EIIB | IPR018113 | Phosphotransferase system EIIB, cysteine phosphorylation site | 404 | 435 | 1.2E-10 |
| ProSiteProfiles | PS51098 | PTS_EIIB type-1 domain profile. | IPR001996 | Phosphotransferase system, IIB component, type 1 | 399 | 481 | 21.993 |
| TIGRFAM | TIGR01998 | PTS-II-BC-nag: PTS system, N-acetylglucosamine-specific IIBC component | IPR010974 | Phosphotransferase system, N-acetylglucosamine-specific IIBC component | 9 | 472 | 6.9E-183 |
| SUPERFAMILY | SSF55604 | IPR036878 | Glucose permease domain IIB | 402 | 474 | 1.24E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.